Introduction
Detecting the adulteration of processed meat with unwanted food ingredients is one of the most important food quality-related issues. Food ingredient authentication is related to human health since ingredients may include allergenic or toxic substances. Furthermore, certain groups of people will not eat specific meats because of their religious food ethics and preferences (Ortea et al., 2012). Hsieh et al. (1997) reported that multispecies adulteration were found in commercial meat products. In addition, products labeled beef only are often intentionally adulterated with pork owing to the economic advantage that pork provides. This is found to be especially in the countries having expensive beef such as Korea, Japan, China, and so on (Singh and Neelam, 2011; Soares et al., 2013).
Food labeling regulations in many countries require that the meat species used in processed meat products must be declared for consumers because of religious food ethics, medical purposes, and personal food preferences (Doosti et al., 2014). However, processed meat products are still mislabeled for meat species, especially pork, which is either intentional or accidental (Tanabe et al., 2007). An accurate detection method for pork therefore needs to be developed for the prevention of food adulteration.
Existing detection methods for the identification of pork in processed meat products rely on protein or DNA analysis. Protein-based analytical methods include immunological assays (Anguita et al., 1996; Chen and Hsieh, 2000), chromatography (Chou et al., 2007), and peptide examination (Aristoy and Toldrá, 2004). However, protein-based analytical methods may not be appropriate for processed meat products since proteins can be denatured during processing. On the other hand, DNA-based analyses such as the polymerase chain reaction (PCR), real time PCR, PCR-restricted fragment length polymorphism (PCR-RFLP), and species-specific PCR are used more frequently in identifying fraudulent meat products (Man et al., 2007; Murugaiah et al., 2009; Soares et al., 2010), since these methods can detect small amounts of DNA and amplify specific target regions (Saiki et al., 1988). DNA-based analyses also have numerous advantages including simplicity, rapidity, sensibility, and specificity (Lockley and Bardsley, 2000).
The objective of this study was to design primers for the PCR amplification of a specific region of pig mitochondrial DNA and subsequently use these primers to identify mislabeled processed meat products.
Materials and Methods
Pig, cattle, chicken, and sheep mitochondrial DNA sequences were obtained from GenBank (database accession no. AF034253, V00654, AY235570, and AF010406, respectively) and aligned using the MUSCLE multiple sequence alignment program (http://www.ebi.ac.uk/Tools/msa/muscle/). The regions most specific and unique to pork were identified among the aligned sequences using BLAST, and primers were subsequently designed. Sequences of the final selected primers were as follows: Pork-Forward (F) (5’- GGT TCT TAC TTC AGG ACC ATC-3’), and Pork-Reverse (R) (5’- GTG TAC GCA CGT GTA TGT AC-3’) (Table 1).
| Primer | Sequence (5’ → 3’) | Length of base pairs (bp) | Length of PCR product (bp) |
|---|---|---|---|
| Pork-F | GGT TCT TAC TTC AGG ACC ATC | 21 | 294 |
| Pork-R | GTG TAC GCA CGT GTA TGT AC | 20 |
To evaluate primer sensitivity, ground meat samples were purchased from butcher’s shop, and processed meat samples were prepared by mixing ground pork (boston butt), ground beef (shank), and ground chicken (breast) as follows: i) 0% pork-50% beef-50% chicken, ii) 1% pork-49.5% beef-49.5% chicken, iii) 2% pork-49% beef-49% chicken, iv) 5% pork-47.5% beef-47.5% chicken, v) 10% pork-45% beef-45% chicken, and vi) 100% pork-0% beef-0% chicken. All mixtures were kneaded by hands, subsequently placed into Eppendorf tube, and cooked in a water bath at 70°C for 3 min. The cooked meat was minced with a knife, and 200 mg was used for DNA extraction.
A total of 35 packaged meat products, such as 14 patties, 8 nuggets, 8 meatballs, and 5 sausages were purchased from a commercial vendor in Itaewon, Korea. Most products were frozen and some products were refrigerated. The product labels indicated that purchase products were manufactured with only chicken, beef, or mixture of these two meats. All products were stored at −20°C until used in the experiments.
Genomic DNA was extracted from 200 mg of each of the processed meat products using the PowerPrepTM DNA Extraction from Food and Feed Kit (Kogenbiotech Co., Ltd., Korea). Six hundred microliters of lysis buffer A and 40 μL of lysis buffer B were added to 200 mg of each of the processed meat products, followed by incubation at 65°C for 1 h. Thereafter, 400 μL chloroform was added and centrifuged at 12,000 rpm for 10 min. Two hundred microliters of the supernatant was mixed with 200 μL binding buffer and 200 μL isopropanol, and the mixture was transferred into the DNA binding column. The DNA was washed twice with 75% ethanol and finally the DNA was eluted by 100 μL distilled water.
The PCR amplification was performed in a 50 μL reaction volume using 25 μL Taq PCR Master Mix Kit (Qiagen, Germany), 2 μL of each primer, 2 μL extracted DNA, and 19 μL of distilled water. The amplification profile was an initial denaturation step at 95°C for 5 min, followed by 40 cycles at 95°C for 20 s, 55°C for 20 s, and 72°C for 30 s. In order to confirm amplification of the target sequence, the PCR product was electrophoresed on a 2% agarose gel in 1×TAE buffer (Biosesang, Korea) at 100 V for 20 min. A 100-bp DNA Ladder (Dynebio Inc., Korea) was used as the size marker and images were captured in a UV-transilluminator (TF-20M, Vilber Lourmat, France).
Results and Discussion
Food adulteration has been an issue for many years in processed meat products. In particular, pork is often mixed in other meat products such as beef, because it is cheaper. Thus, various pork detection methods have been developed. Among these methods, species-specific oligo Vilber Lourmat nucleotide primers have been used to detect pork in processed meat products. This study targeted the 294-bp long pig mitochondrial DNA D-loop region since mitochondrial DNA is highly conserved in many animal species, is stable from heating, and can be used to detect pork fat (Montiel-Sosa et al., 2000). This suggests that the primers designed in our study could be used to detect pork in heat-treated products.
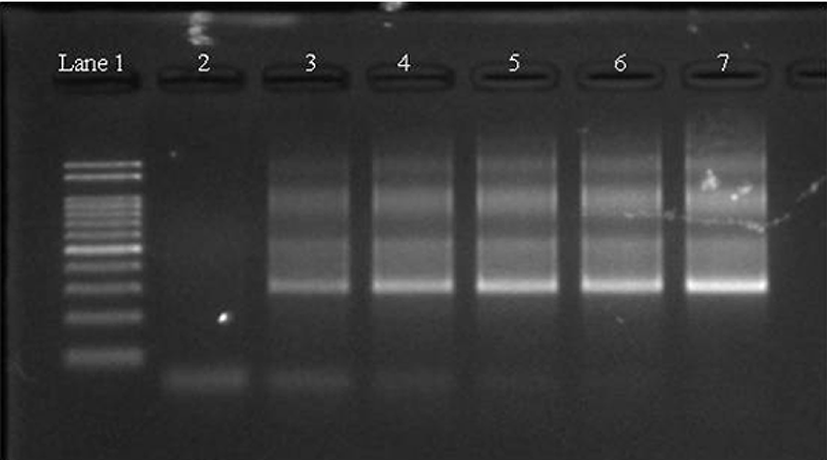
There are two methods to detect pork adulteration in processed meat products, namely protein and DNA analysis. DNA-based methods are used more frequently than protein-based methods to detect adulterated food, because of the limitations of protein-based methods such as protein denaturation following heating (Fajardo et al., 2010). Meat mixtures were prepared with the inclusion of pork at a concentration of 0%, 1%, 2%, 5%, 10%, and 100%, and subsequently heated to determine primer compatibility and detection limits. Study results indicate the developed primers could detect as little as 1% pork in the heat treated meat products (Fig. 1).
To monitor pork adulteration in commercial processed meats with the primers developed in our study, 35 processed meat products were purchased including patties, nuggets, meatballs, and sausages labeled as 0% pork. Of the 35 products, three (8.6%) were pork-positive (Table 2). The positive samples were one meatball and two sausage products. The meatball was labeled as beef only, and the sausages were labeled as beef and chicken, and chicken only. Murugaiah et al. (2009) reported that adulteration occurs predominantly in comminute or ground meat products. In our study, all pork-positive samples were from ground meat products.
Most ground meat is accidentally adulterated with unwanted meat species during the process of grinding. It is, thus, essential that grinders are thoroughly cleaned when changing meat species, alternatively different meat grinders should be used for different meat species. However, pork is also intentionally added to beef products because of production costs (Doosti et al., 2014). In many countries, labels declaring the specific meat species used are mandatory because of food allergens, religious food ethics, and prevention of food fraud (Ayaz et al., 2006; Gendel, 2012; Soares et al., 2013); however, as shown in Table 2, adulteration still occurs in commercial products. It is therefore important that an accurate and simple method to detect pork adulteration in processed meat products is developed.
In conclusion, the primers developed in our study could be used to detect the presence of pork adulteration in various processed meat products, even when heat treated. In addition, pork adulterated commercial meat products were identified, and thus, government agencies should consider monitoring mislabeling of meat products that include pork.













