Introduction
Campylobacteriosis is an infectious disease that can cause gastrointestinal symptoms including diarrhea, abdominal pain, and vomiting. Guillaine-Barre syndrome is one of the complications of campylobacteriosis that can damage nerve system of humans (Yang et al., 2013). The European Center for Disease Prevention and Control and the European Food Safety Authority ranked campylobacteriosis as the most common zoonosis in Europe (Eurosurveillance Editorial Team, 2012). The most common species that can cause campylobacteriosis in humans is Campylobacter jejuni, followed by Campylobacter coli (Tam et al., 2003). Contact with live animals and consuming of raw poultry have been defined as the main sources of Campylobacter infection (Tam et al., 2003).
To confirm species origin for pathogens, analytical methods are mostly based on protein or DNA analysis (Ayaz et al., 2006). However, protein-based isolation method has a weakness. When the sample is exposed to high temperatures and pressure, the protein has a general tendency to degenerate (Murugaiah et al., 2009). Compared with protein-based methods, DNA based methods are more dependable because of their unique variability and high stability. Among DNA based methods, polymerase chain reaction (PCR) methods, including conventional PCR, real-time PCR (Cammà et al., 2012; Kesmen et al., 2012), and PCR-RFLP (Chen et al., 2010; Haider et al., 2012) have high specificity and accuracy. Recently, specific nanoparticles, including silver (AgNPs) and gold (AuNPs) nanoparticles, have been widely used as components of new technologies to detect pathogens (Du et al., 2013), hazardous materials (Li et al., 2009), DNA (Benedetto et al., 2011), small molecules (Lv et al., 2013), and aptamers (Ping et al., 2012). Aptamer-based detection has been carried out in our previous research (Kim et al., 2018). When various solutions change from dispersion to aggregation state, specific nanoparticles exhibit significant color change because of their unique optical properties, that can be observed using a UV-visible wave-based spectrophotometer or the naked eye (Du et al., 2020; Shams et al., 2019). Thus, this research was designed to use AuNPs. A few types of AuNP-based colorimetric assays have been previously reported. In particular, the colorimetric assay based on thiol-labeled PCR primer and AuNPs has been previously reported for the detection or diagnosis of disease (Htoo et al., 2019; Osmani Bojd et al., 2017). The nanoparticle method is favored in the surveillance of pathogens over other PCR methods, due to its advantages over other PCR methods such as conveniency and time saving. To our knowledge, however, few studies have focused on the selective detection of fastidious foodborne pathogens such as Campylobacter. Since conventional ways to detect Campylobacter based on culture method require more than approximately 4 days (Masdor et al., 2016), various rapid detection methods have been developed for decades (Yang et al., 2013).
In this study, we aimed to develop a colorimetric assay integrating PCR and AuNP conjugation technology. The current study also outlined a colorimetric assay for the PCR-amplified Campylobacter genes, that could be directly identified by the naked eye.
Materials and Methods
All oligonucleotides (Denis et al., 1999) were synthesized and purified by Bionics (Seoul, Korea; Table 1). For the inclusivity test, some C. jejuni (A total of four C. jejuni strains: ATCC 33560 and 3 wild type strains from chicken carcasses) and C. coli strains (A total of four C. coli strains: ATCC 33559 and 3 wild type strains from chicken carcasses) were used. Campylobacter strains were isolated from chicken carcasses from a local slaughterhouse and a poultry farm. Non-Campylobacter strains for the exclusivity test including Salmonella Enteritidis and Escherichia coli, were also isolated from the collected chicken carcasses.
| Target gene | Sequence (5′-3′) | Amplicon size | Reference |
|---|---|---|---|
| mapA (Campylobacter jejuni) forward | SH-C12-CTA TTT TAT TTT TGA GTG CTT GTG | 589 bp | (Denis et al., 1999) |
| mapA (C. jejuni) reverse | GCT TTA TTT GCC ATT TGT TTT ATT A | ||
| ceuE (Campylobacter coli) forward | SH-C12-AAT TGA AAA TTG CTC CAA CTA TG | 462 bp | (Denis et al., 1999) |
| ceuE (C. coli) reverse | TGA TTT TAT TAT TTG TAG CAG CG |
All Campylobacter strains were incubated in Bolton broth (Oxoid, Hampshire, UK) at 42°C for 42 h under microaerobic condition (5% O2, 10% CO2, and 85% N2), while non-Campylobacter strains were incubated in tryptic soy broth (Oxoid) at 37°C for 24 h. Additionally, phosphate-buffered saline was applied to dilute samples in the sensitivity test of AuNP-PCR and 4 Log CFU of Campylobacter and non-Campylobacter strains were incubated in 30 g of chicken meat samples with 30 mL of Bolton broth (42°C for 42 h) and tryptic soy broth (37°C for 24 h). Thereafter, 10 μL of cultured bacteria was used for AuNP-PCR direct detection test.
Approximately 1 mL cultures were used for DNA extraction. DNA was extracted by using QIAamp minikit (Qiagen, Hilden, Germany). The concentration of extracted DNA was measured by using Nanodrop 2000 (Thermo Scientific, Wilmington, DE, USA) and genomic DNA was stored at 4°C before PCR amplification.
PCR was conducted under the following conditions for the amplification of the mapA (C. jejuni) and ceuE (C. coli) genes. The reaction was carried out in a 50 μL mixture with 50 mM KCl, 1.5 mM MgCl2, 10 mM Tris-HCl, 0.5 mM of deoxynucleoside triphosphate, 200 nM of forward/backward primer, 1 unit of Taq DNA polymerase (TaKaRa, Kusatsu, Japan), and 2 μL (200 ng) of template DNA. The details of primers used in this study is presented in Table 1. The amplification initiated with 10 min at 95°C, followed by 30 cycles of 30 s at 94°C, 90 s at 59°C, and 60 s at 72°C, and then 10 min at 72°C. PCR was conducted in a Veriti 96 well thermal cycler (Applied Biosystems, Waltham, MA, USA). Exactly 2 μL of PCR products were then used for a gel electrophoresis with 1.0% agarose gel. The samples were run at 100 V for 30 min, followed by some amount of PCR products being visualized with image analysis software (image lab software V3, Bio-Rad Laboratories, Hercules, CA, USA). The length of expected PCR product was 589 bp and 462 bp for mapA and ceuE gene, respectively.
Sodium citrate dehydrate and HAuCl4 were from Sigma Aldrich (St. Louis, MO, USA). AuNPs (13 nm in diameter) were prepared via HAuCl4 citrate reduction. All glassware was cleaned in KOH-IPA (potassium hydroxide+isopropyl alcohol) solution and washed with pure H2O. A 50 mL of a 38.8 mM trisodium citrate solution was rapidly mixed with HAuCl4 solution (1 mM, 500 mL), resulting in a change of color from yellow to dark red. The solution was then refluxed for another 15 min to cool it down to 25°C. The characteristics of the AuNPs were verified using a surface plasmon band centered at 520 nm (Grabar et al., 1995).
The center of the surface plasmon band of AuNPs was 520 nm. A 6 μL of PCR product was mixed with 40 μL of the gold colloid for the colorimetric assay. They were mixed for 1 min of mixing, and another 10 μL of 1 M NaCl was added. The color was identified by the naked eye, and they were quantified using Nanodrop 2000 (Thermo Scientific). Since the aggregations between AuNPs at 520 and at 650 nm are different, the ratio of absorbance 520/650 (Abs520/Abs650) was chosen.
The colloidal gold solution was dropped onto 1,000 mesh of copper grids that was coated by carbon, followed by being dried in ambient conditions. Images were acquired using TEM JEM-2100 (JEOL, Tokyo, Japan) from Seoul National University. At least ten locations on the TEM copper grid were examined.
Analysis of different Abs520/Abs650 data for Campylobacter spp. for specificity was compared to that for non-Campylobacter spp. and the data for various concentrations of PCR products in the sensitivity analysis were analyzed using one-way ANOVA, which was conducted using GraphPad Prism software (GraphPad Software, San Diego, CA, USA).
Results and Discussion
Fig. 1 represents the main principle of the suggested method. The main idea behind the AuNP colorimetric assay was the dispersion and aggregation of colloidal AuNPs. It was effectively demonstrated that the interparticle movement and pulsive forces are the main reason for AuNP stabilization or aggregation (Napper, 1983; Zhao et al., 2008). AuNP electrostatic stabilization relies on the surface charge of the repulsive electric double layer. The layer can stabilize the colloids against Van der Waals forces (Evans and Wennerström, 1999). When the electric double layer of AuNP is suppressed, the electrostatic repulsion force dramatically decreases at high concentrations of salt. That is why citrate-combined AuNPs are stabilized in water but aggregated in salty conditions (Zhao et al., 2008). Thus, the covalently combined complex of thiolated double-strand DNA (dsDNA) and monodispersed AuNP can resist the aggregation regardless of salt addition.
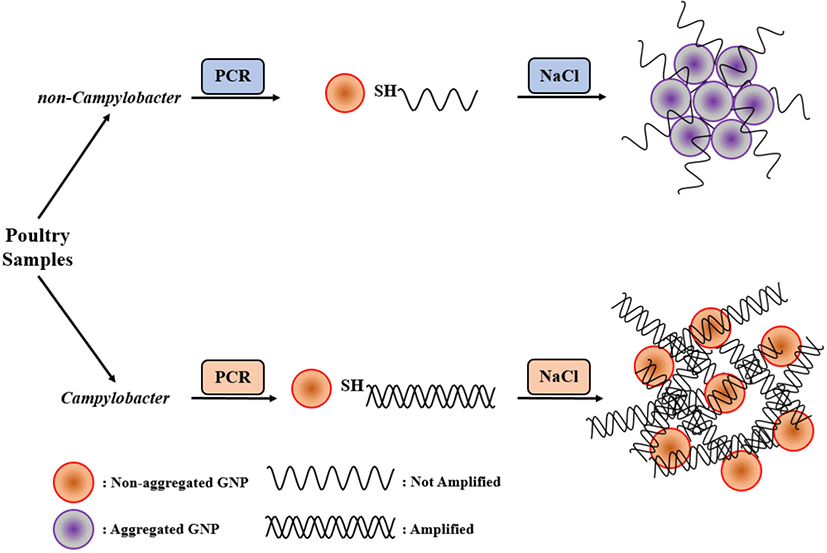
AuNPs maintain a red color in stable state. When they aggregated, the state of AuNPs becomes unstable, causing change of color from red to blue or purple. The main focus of the current research was that AuNPs can be modified by thiolated PCR products, that prevented AuNPs from salt-induced aggregation (Chan et al., 2014; Fu et al., 2013; Jyoti et al., 2010). Because of this attribute, we modified unique Campylobacter target primers with an unlabeled backward primer and a thiol-labeled forward primer. Therefore, thiol-labeled target gene sequence was obtained. When AuNPs were mixed with thiolated PCR products, they were surrounded by a thick layer that consisted of dsDNA. Therefore, this structure tended to make a change of color.
For a successful result, AuNPs need a thick obstacle with a lot of negative charges to inhibit their aggregation. We modified the primer for mapA of C. jejuni (589 bp) and ceuE of C. coli (462 bp) for PCR amplification. mapA, which is usually called membrane-associated protein A gene, does not cross-react with C. coli proteins (Gonzalez et al., 1997; Stucki et al., 1995). ceuE is a lipoprotein-encoding gene which has been used for C. coli identification (Gonzalez et al., 1997). To confirm PCR efficiency, thiol-labeled PCR primers used for C. jejuni and C. coli were the same as unlabeled primers used for the conventional PCR.
PCR and gel electrophoresis were conducted to analyze the PCR products. Fig. 2 shows that PCR products from thiol-labeled and unlabeled primers are identical in size. It shows that the result obtained using the thiolated primers did not show any size difference to conventional PCR products.
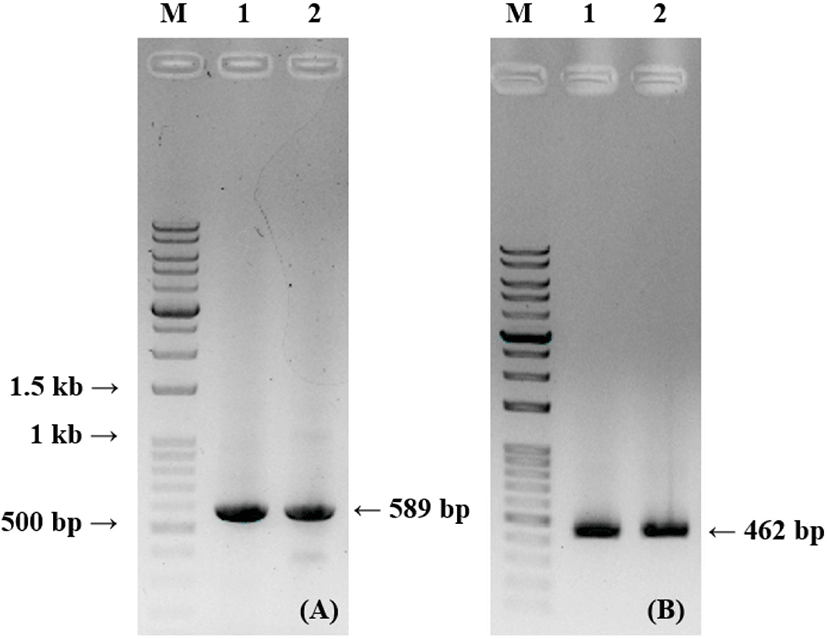
For the detection of Campylobacter spp., thiol-labeled PCR products prevented AuNP aggregation based on salt-inducing. Both unlabeled and thiol-labeled PCR products were added into the AuNP solution and the color change was compared. After adding NaCl solution, AuNP solution with unlabeled PCR products turned from red to blue and that with thiol-labeled products remained red. Spectroscopic analysis verified a sharp peak of surface plasmon resonance (SPR) at approximately 520 nm. This narrow SPR peak appeared due to the red colored solution containing thiol-labeled PCR products after salt adjusting. A broad SPR band (530–650 nm) seemed in the blue colored solution with unlabeled PCR products, indicating AuNP aggregation after salt addition (Fig. 3). To confirm the characteristic features of AuNPs in the colorimetric assay, different behaviors of the salt-induced aggregation of AuNPs were observed using TEM JEM-2100 (JEOL; Fig. 4).
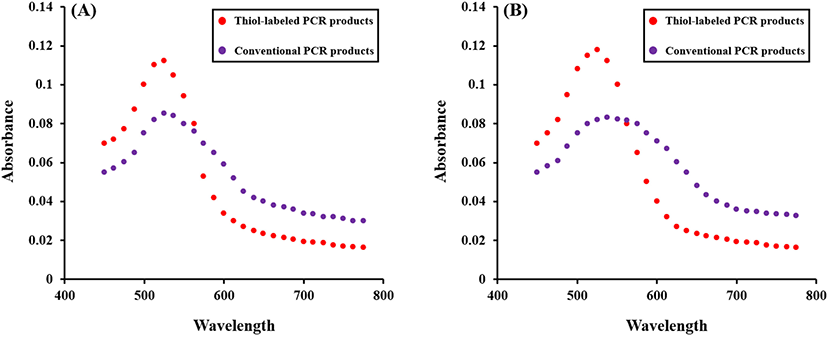
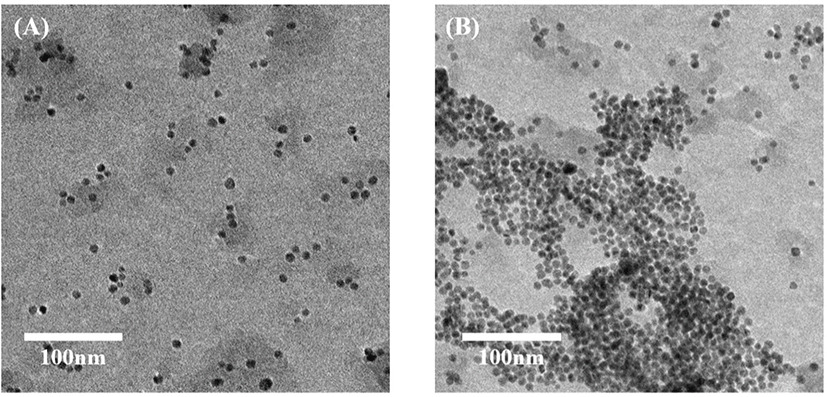
A color change was observed in the AuNP solution containing non-labeled PCR products, whereas the AuNP solution with thiol-labeled PCR products are still red (Fig. 5). This can be identified with the naked eye.
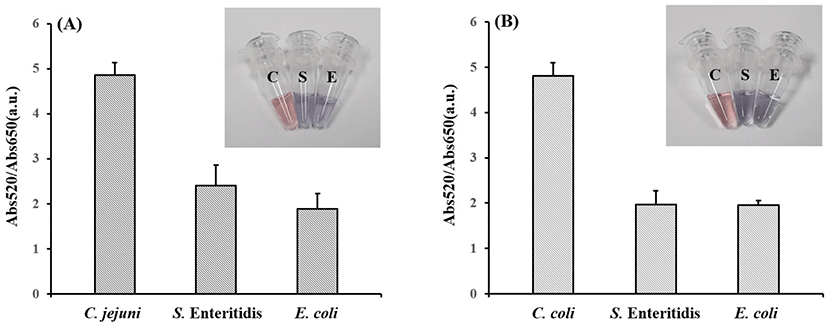
Abs520/Abs650 of C. jejuni and C. coli isolates were significantly different (C. jejuni: 4.86±0.27; C. coli: 4.81±0.28) compared to that of S. Enteritidis and E. coli (C. jejuni p-value: 0.0085; C. coli p-value: 0.0239) in five replicates. Thus, 200 ng/μL of mapA and ceuE were detected in the sample with thiol-labeled products after salt adjusting.
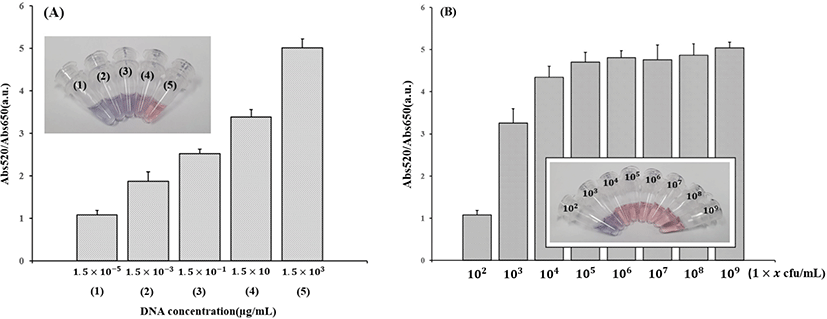
For the determination of the sensitivity, different concentrations of mapA from C. jejuni were tested. Abs520/Abs650 of the detected samples are shown in Fig. 6. Fig. 6A shows that at least 15 ng/μL of C. jejuni mapA PCR product was needed to detect C. jejuni using the optical method. At this concentration, Abs520/Abs650 was 3.38±0.17 (mean±SD). Fig. 6B shows that at least 1×103 CFU/mL of C. jejuni was needed to use the colorimetric method. At this concentration, Abs520/Abs650 was 3.26±0.35 (mean±SD). These results represented that the Abs520/Abs650 increased with the increasing number of PCR products and cells. To verify the precision of the assay, one-way ANOVA was used to determine the p-value after measuring the absorption at different concentrations in five replicates.
More PCR products can be obtained by applying more amplification cycles. However, too many amplification cycles lead to low efficiency because of the formation of non-specific bands due to DNA degeneration (Cha and Thilly, 1993). Therefore, 30 cycles of PCR protocol was selected in this study. The colorimetric system was controled by NaCl. The concentration of NaCl affects the dynamics of the method as well as the sensitivity of the detection. The limit of detection of various types of detection methods are compared in Table 2. qPCR is slightly more sensitive than AuNP-PCR (Table 2). However, the detection limit of the AuNP-PCR was similar or superior to other detection methods such as ELISA, AuNP-adjusted aptasensor, and Lateral flow (Table 2).
| Methods | Target bacteria | LOD (CFU/mL) | Reference |
|---|---|---|---|
| ELISA | Campylobacter jejuni | 4.5×104 | (Che et al., 2001) |
| AuNP-adjusted aptasensor | C. jejuni | 7.2×105 | (Kim et al., 2018) |
| QD adjusted fluorescent biosensor | C. jejuni | 1×103 | (He et al., 2018) |
| Lateral flow | C. jejuni | 1×107 | (Wadl et al., 2009) |
| qPCR | C. jejuni | 1.25×102 | (Suh et al., 2014) |
| AuNP-PCR assay | C. jejuni | 1×103 | The present study |
As we stated above, the thiol-labeled PCR primer and AuNPs have been applied in many research fields for the detection or diagnosis of disease. Htoo et al. (2019) used thiol-labeled PCR primer and unmodified AuNPs for the detection of target RNA for the diagnosis of prostate cancer and found that the assay was specific and sensitive for the RNA detection in prostate cancer cell lines with a visual detection limit of 31.25 ng/reaction. Osmani Bojd et al. (2017) reported application of thiolated AuNP probes and multiplex PCR for the detection of Staphylococcus epidermidis. They found that the minimum quantity of target DNA for multiplex PCR was 1 ng/mL and for color and absorption altercation of solution in colorimetric assay was 20 ng/mL. Since Campylobacter is fastidious bacteria that requires long incubation time and labors, development of this types of rapid and convenient isolation method is highly useful for the screening of the pathogen. In the present study, the newly developed method showed slightly lower sensitivity compared to qPCR according to Table 2. However, this method could be favored over qPCR especially in field tests as the assay could be conducted without further steps that require cost, labor, and detection device. With the assay, the positives could be easily distinguished from the negatives with the naked eye. Thus, the developed assay could be finished only with thermal cycler and optical detection, while conventional or qPCR still need further steps after PCR cycles. Even though qPCR is superior to this assay in terms of sensitivity, the developed method could be widely used in many research fields considering its convenience for the detection of pathogens.
To estimate the detection ability of AuNP-PCR in food pathogenic bacteria, the method was applied to a food sample. Three groups of chicken meat samples were purchased from local retails. A group was artificially spiked with an overnight culture of C. jejuni, while the others were contaminated with S. Enteritidis and E. coli. Artificially contaminated samples were cultured at 42°C overnight and 10 μL of each sample was directly adjusted to the AuNP-PCR assay. The results of these assays are shown in Fig. 7. When thiol-labeled primers were used in the rinsate of fresh chicken meat, the color of the AuNP solution changed from red to blue after salt adjusting. Abs520/Abs650 was 1.06±0.22 (mean±SD). However, if the C. jejuni template was present, the color of AuNP solution is still red after salt adjusting. This result suggested that the AuNPs remained dispersed, while the Abs520/Abs650 of mapA was 4.30±0.3. With the spectroscopic results, the sample groups can be clearly distinguished.
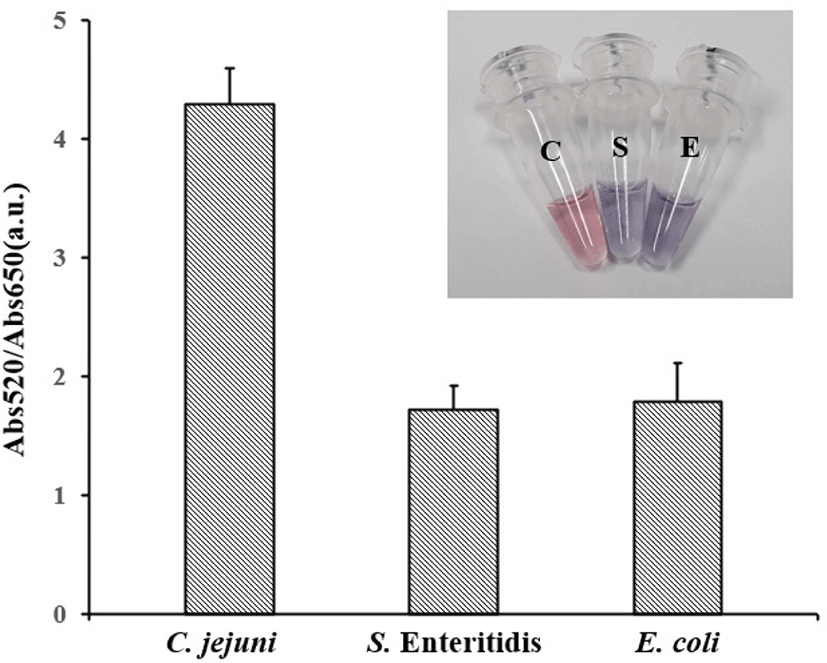
The detection ability of the AuNP-PCR assay for C. jejuni and C. coli was well defined according to sensitivity assays results. The length of PCR amplicons and amplification efficiency were important factors for the detection. Previous research found that AuNPs highly tend to aggregate when the PCR amplicons were shorter than 400 bp. When the amplicons were longer than 400 bp, it became a limiting factor for efficient detection (Fu et al., 2013). As we stated above, AuNPs require a thick barrier with a lot of negative charges to inhibit the aggregation. We selected the 589 bp mapA for C. jejuni detection and the 462 bp ceuE for C. coli detection.
Conclusion
The suggested assay enables the generation of a PCR product with high amplification efficiency and the convenience of a colorimetric assay. Moreover, leading to a limit of optical detection of density around 15 ng/μL of Campylobacter verified high specificity. The suggested assay has a couple of advantages. Above all, the PCR colorimetric method has high sensitivity, specificity, and amplification efficiency. Furthermore, this method avoids bacterial culture steps, reduces the time needed, and requires no complicated apparatus. Moreover, the proposed method allows the detection of pathogens even by the naked eye. Therefore, it appears that this assay has high potential for a various bio-related detections. Lastly, as we stated above, this method does not require complex devices, while detecting conveniently with the naked eye. The current research reported a novel and unique colorimetric assay for the detection of Campylobacter spp. in raw poultry. Considering high specificity of the assay, the accurate identification of Campylobacter spp. from other environmental samples is also available with this method.













