Introduction
Correctly identifying meat species in food products is very important for authenticating food, promoting food safety, and preventing food adulteration. Meat species identification is a critical issue because of the different forms of meat adulteration, including the replacement of expensive meat with cheaper meat, the presence of less meat than indicated on the product label, and the inclusion of meat in non-meat (vegetarian) products (Zia et al., 2020). Furthermore, accurate meat species identification is important for satisfying religious requirements for certain foods (e.g., Halal meat). Islamic law strictly forbids the consumption of some meat products, especially pork. Thus, there are authenticity problem and religious reason for protecting consumers by detecting pork in food products.
Among the various analytical methods available for detecting meat species in foods, highly sensitive and specific DNA-based methods have been commonly applied (El Sheikha et al., 2017). PCR-based methods involving random amplified polymorphic DNA (Arslan et al., 2005; Mane et al., 2008), restriction fragment length polymorphisms (Hossain et al., 2016; Rahmati et al., 2016), DNA barcoding (Kane and Hellberg 2016; Naaum et al., 2018), and real-time PCR (Amaral et al., 2017; Mohamad et al., 2018) are frequently used because they enable rapid and precise detection of meat species. Among these options, real-time PCR with a species-specific primer and a TaqMan probe is the most suitable and widely used method for identifying meat species (Ali et al., 2012; Kim et al., 2016). Furthermore, there are numerous available commercial master mixes and platforms, with increasing interest in the TaqMan real-time PCR assay.
Several previous studies investigated the effects of different master mixes and platforms on real-time PCR performance characteristics. For example, Sohni et al. (2008) evaluated five commercial real-time PCR reagents used for detecting Bacillus anthracis by comparing their PCR efficiencies and limits of detection (LODs). Stephens et al. (2010) compared five master mixes used for detecting the Ebola virus regarding their performance characteristics such as sensitivity and PCR efficiency. Meanwhile, Buzard et al. (2012) conducted a multi-platform comparison of nine commercial master mixes used for detecting bioterrorism agents. Furthermore, Eischeid and Kasko (2015) compared the utility of four master mixes for identifying a shrimp allergen in a real-time PCR assay. All of these studies emphasized the importance of choosing a suitable master mix and platform, both of which can influence the sensitivity and efficiency of a PCR assay. However, there have been relatively few comparative studies regarding real-time PCR master mixes and platforms used for identifying meats, especially pork. Moreover, most of these studies did not test whether the PCR assay was specific for meat from a particular animal species.
Therefore, the main objective of this study was to optimize a real-time PCR assay for detecting porcine material. Seven commercial master mixes were compared regarding specific performance criteria, including LOD, PCR efficiency, specificity, total cost, and time. The reliability of the data was confirmed with two real-time PCR platforms.
Materials and Methods
The following seven commercial TaqMan master mixes were evaluated: TaqMan Universal PCR Master Mix (Applied Biosystems, Waltham, MA, USA), MG 2X qPCR MasterMix (TaqMan) with ROX (CancerROP, Seoul, Korea), Express qPCR Supermix Universal (Invitrogen, Waltham, MA, USA), PowerAmp Real-time PCR Master Mix II (Kogene Biotech, Seoul, Korea), Luna Universal Probe qPCR Master Mix (New England Biolabs, Ipswich, MA, USA), QuantiNova Probe PCR Kit (Qiagen, Hilden, Germany), and Premix Ex Taq (Probe qPCR), ROX plus (Takara, Shiga, Japan).
Four raw meat samples including pig (Sus scrofa domesticus), dog (Canis familiaris), chicken (Gallus gallus), and cow (Bos taurus) were purchased from local supermarkets in Korea, and human cheek cells were obtained after rinsing the mouth with 1 mL 8% NaCl.
Meat-processed foods including 4 types of pork-containing products (dumpling, ham, pork cutlet, and sausage), 3 types of beef-containing products (beef curry, beef stock, and meatballs), and 3 types of chicken-containing products (chicken teriyaki, chicken sausage, and chicken stock) were purchased from local markets in Korea.
Genomic DNA was extracted from 200 mg of finely ground samples. For all samples, DNA was extracted with the PowerPrep™ DNA Extraction from Food and Feed Kit (Kogene Biotech) according to the manufacturer’s instruction. The concentration and quality of extracted DNA were determined by Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The DNA concentration of all samples was adjusted to 50 ng/μL and a 10-fold serial dilution series of porcine DNA was prepared to generate a real-time PCR standard curve.
The porcine-specific primer set [5′-CGCCTCACTCACATTAACCA-3′ (forward) and 5′-AAGGGGACTAGGCTGAGA GTG-3′ (reverse)] and TaqMan probe [5′-FAM-CACTGACTATTCTAACCATCCCAA-BHQ1-3′] were designed as follows. Various porcine DNA sequences from the National Center for Biotechnology Information GenBank database (http://www.ncbi.nlm.nih.gov/Genbank) were aligned with the ClustalW program (http://www.ebi.ac.uk/clustalw/). The primer set was designed to specifically target the conserved region of the ND5 gene (NADH dehydrogenase subunit 5) from the porcine mitochondrial genome to produce a 141-bp amplicon. The TaqMan probe was tag with FAM and BHQ1 at the 5′ and 3′ ends, respectively. The primer set and probe were synthesized by Bioneer (Deajeon, Korea).
The reaction mixture for the real-time PCR assay comprised 10 μL each master mix, 500 nM primer set, 500 nM TaqMan probe, 1 μL DNA (10-fold serial dilution series), and distilled water for a final volume of 20 μL. The manufacturer’s recommended thermal cycling conditions used in this study are listed in Table 1. To determine the LODs and PCR efficiencies, all samples of the 10-fold serial dilution series were analyzed in triplicate on the same 96-well optical reaction plate (Applied Biosystems). The real-time PCR assay was completed using the StepOnePlusTM Real-Time PCR system (Applied Biosystems) and the CFX ConnectTM Real-Time PCR System (Bio-Rad, Hercules, CA, USA).
All data were analyzed with StepOnePlusTM Software (version 2.3) (Applied Biosystems) and CFX ManagerTM Software (Bio-Rad). Moreover, these programs automatically generated the standard curve and determined the PCR efficiency, which was calculated as E=−1+10(−1/slope).
Results and Discussion
To compare the LODs and PCR efficiencies, 10-fold serial dilutions of porcine DNA (0.0005–50 ng/μL) were analyzed in a real-time PCR assay involving seven master mixes and two platforms (Table 2). All master mixes performed stably on both platforms, but the LODs of the master mixes ranged from 0.5 to 5 pg/reaction (rxn). Sensitive and reliable amplifications were observed for the Kogene Biotech, Invitrogen, Qiagen, and New England Biolabs master mixes, with an LOD of 0.5 pg/rxn on both platforms. In contrast, the LOD for the Applied Biosystems and CancerROP master mixes differed depending on the platform. Specifically, the LOD for the Applied Biosystems master mix was 10 times lower on the StepOnePlus platform (0.5 pg/rxn) than on the CFX Connect platform (5 pg/rxn). Conversely, the LOD of the CancerROP master mix was 10 times lower on the CFX Connect platform (0.5 pg/rxn) than on the StepOnePlus platform (5 pg/rxn). The Takara master mix was the least sensitive, with an LOD of 5 pg/rxn. The PCR efficiencies of the seven master mixes ranged from 84.96% to 108.80% depending on the master mixes and platforms (Fig. 1). The Kogene Biotech master mix on the CFX Connect platform performed best, with an efficiency of 100.49% (correlation coefficient, r2=0.9997). Meanwhile, the worst performance was observed for the Applied Biosystems master mix on the CFX Connect platform, with an efficiency of 84.96% (correlation coefficient, r2=0.9943).
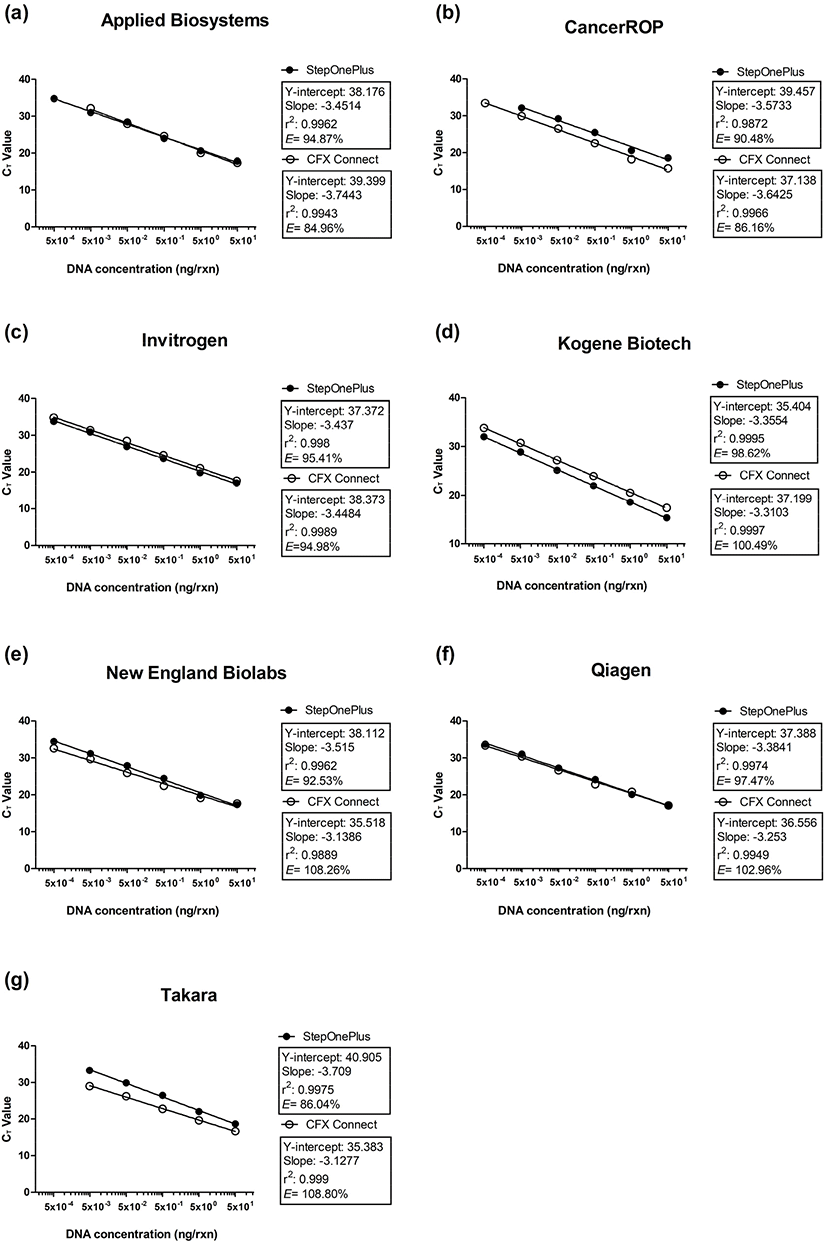
The data presented herein revealed that the sensitivity and efficiency of a real-time PCR assay varied depending on the master mix and platform used. The observed differences were due to the DNA polymerase and buffer in the master mix, both of which influence the amplification efficiency and ability to detect specific DNA sequences (Wolffs et al., 2004). Previous studies that evaluated the utility of DNA polymerases for amplifying DNA samples collected during forensic analyses compared nine DNA polymerases (AmpliTaq Gold, Bio-X-Act Short, ExTaq Hot Start, KAPA2G Robust, OmniTaq, PicoMaxx High Fidelity, rTth, Taq, and Tth). The resulting LODs and PCR efficiencies differed depending on the DNA polymerase used (Hedman et al., 2009; Hedman et al., 2010). Other studies demonstrated that the master mix buffer components, such as Mg2+ and bovine serum albumin, influence DNA polymerase activity (Bustin, 2004; Kreader, 1996). Therefore, the variability in the LODs and PCR efficiencies in the current study was because the seven tested master mixes comprise a different DNA polymerase–buffer system. Moreover, the suitability of a particular DNA polymerase–buffer system may depend on the DNA target.
This comparative study of seven master mixes may be useful for the development of a very sensitive real-time PCR assay for detecting porcine material. A comparison with several previous real-time PCR-based studies of porcine detection indicated that the LOD (0.0005 ng/rxn) of this study is lower than that (0.01 ng/rxn) of a previous study by Rodríguez et al. (2005), but is similar to the LOD (0.0001 ng/rxn) of a study by Kesmen et al. (2009), which involved the most sensitive real-time PCR system for detecting porcine material. Additionally, appropriate PCR efficiencies were reportedly between 90% and 110% (Adams, 2006). In this study, the PCR efficiencies of most of the master mix and platform combinations were between 84.96% and 108.80%. Moreover, the highest efficiency (100.49%) observed for the Kogene Biotech master mix on the CFX Connect platform was greater than the previously reported PCR efficiencies of 75.83% (Rodríguez et al., 2005), 91.57% (Sakai et al., 2011), and 103.98% (Kesmen et al., 2009).
The specificity of the seven master mixes was tested with 50 ng/μL DNA extracted from four animal species (human, dog, cow, and chicken), with porcine DNA used as a positive control. All samples were analyzed in triplicate along with a no-template control, and the results are presented in Table 3. The positive control was amplified in all seven master mixes, with CT values between 15.35±0.37 and 18.69±0.37. A lack of non-specific amplification was observed for the Applied Biosystems, Takara, and Kogene Biotech master mixes on both platforms. Meanwhile, non-specific amplifications were detected for the Invitrogen and New England Biolabs master mixes, but only on the CFX Connect platform, while they were observed for the Qiagen and CancerROP master mixes on both platforms.
An additional sequence alignment analysis of five animal species (pig, human, dog, cow, and chicken) with the ClustalW program was used to verify the specificity of the primer sets and probes. The primer and probe sequences were completely complementary to the mitochondrial DNA of Sus scrofa domesticus (NC012096.1, AF486858.1, AF486866.1, AY574046.1, DQ518915.2, EU117375.1, KJ746666.1, and KC469587.1), but not to the sequences from other species, specifically Homo sapiens (GU170821.1), Canis familiaris (AY729880.1), Bos taurus (GU947021.1), and Gallus gallus (KM096864.1). A conventional PCR was applied to confirm the specificity of the primer set developed in this study. The electrophoretic separation of amplicon revealed a lack of non-specific products, confirming that the primer set developed in this study is specific for porcine material (data not shown). However, non-specific amplifications were observed for the real-time PCR assay depending on the master mix and platform. A recent study revealed that non-specific amplifications during real-time PCR assays may occur after 27 cycles depending on the master mix and thermal cycling conditions, which is earlier than the reported threshold of 34 cycles (Ruiz-Villalba et al., 2017). In the current study, non-specific amplifications occurred with CT values ranging from 32.75 to 34.91, which are consistent with the data from an earlier study. However, the reason why certain master mixes result in non-specific amplifications remains unknown.
Following a comparison of seven master mixes and platforms based on the LOD, PCR efficiency, and specificity, we selected PowerAmp Real-time PCR Master Mix II and the CFX Connect platform as the best combination for detecting porcine DNA. Pork is one of the most widely used meats in processed meat products. The presence of pork in processed foods may cause allergic reactions in some sensitive people, and it may be used for food adulteration (Soares et al., 2010; Tanabe et al., 2007). Furthermore, it is banned in halal foods.
The real-time PCR assay developed in this study was used to determine the presence or absence of porcine material in various processed foods. The analysis of 10 commercial processed meat products showed positive reactions for all pork-containing products (dumpling, ham, pork cutlet, and sausage) and negative reactions for all beef-containing products (beef curry, beef stock, and meatballs) and chicken-containing products (chicken teriyaki, chicken sausage, and chicken stock) (Table 4). This result was consistent with the ingredients listed on the label of the commercial products. Therefore, the porcine-specific real-time PCR assay could be used for the identification and detection of hidden allergens and food adulterants in processed foods and the analysis of halal foods.
1) PowerAmp Real-time PCR Master Mix II and the CFX connect platform were used for porcine-specific real-time PCR assay.
The seven master mixes included in this study were compared in terms of their costs and times required for the real-time PCR (Table 5). The cost per reaction of the seven master mixes ranged from US$ 0.33 to 1.26. The most and least expensive master mixes were from Invitrogen and CancerROP, respectively. On the basis of the manufacturer’s recommended protocols, the total real-time PCR run-times ranged from 10.75 to 57.50 min, with the longest and shortest run-times associated with the CancerROP and Qiagen master mixes, respectively.
Conclusion
Seven commercial master mixes were evaluated to optimize the real-time PCR assay conditions for detecting porcine materials. Additionally, LODs, PCR efficiency, and specificity, as well as the total times and costs of each master mix were analyzed using two real-time PCR platforms. Consequently, a sensitive (LOD of 0.5 to 5 pg/rxn) and efficient (84.96% to 108.80%) real-time PCR-based porcine detection system was developed. However, the LODs and PCR efficiencies varied depending on the master mixes and platforms. Moreover, a specificity test involving four animal species unrelated to pig revealed that non-specific amplifications were not observed on both platforms for only three master mixes from Applied Biosystems, Kogene Biotech, and Takara. These results prove that real-time PCR assays can be influenced by the master mixes and platform. The analysis of commercial processed meat products using the porcine-specific real-time PCR assay showed results that were consistent with the ingredients listed on the label. The result of this comparative study may be useful for optimizing porcine detection system based on a real-time PCR assay.













