Introduction
Various studies have demonstrated that wild animals and their surrounding environments are significant reservoirs of antibiotic resistance genes and antibiotic resistant bacteria (Cantas et al., 2013; Costa et al., 2008; Karesh et al., 2012). This is a growing public health issue, due to increased wildlife contact with humans, as well as livestock and domestic animals due to co-habitation on the same farm (van den Honert et al., 2018). In addition, there is a rising trend in the consumption of game meat over recent years (Cantas et al., 2013; Dias et al., 2015). Majority of these wild animals are slaughtered in field abattoirs (van Shalkwyk and Hoffman, 2010). This can potentially increase the risk that bacteria from the gut and skin of animals can cross-contaminate meat during the slaughter process, typically due to inadequate hygiene conditions and handling (Bakhtiary et al., 2016; van Shalkwyk and Hoffman, 2010). Schlegelová et al. (2004) has stated that bacteria which contaminate meat can be the source of foodborne diseases and also possibly a cause of drug resistant human pathogenic bacteria (Schlegelová et al., 2004).
Although studies are limited, especially on bacteria from wildlife, several studies have been conducted on the antibiotic resistance patterns within a meat processing facility and conclude that cross-contamination of antibiotic resistant bacteria onto the meat does occur and originates from various sources such as the animal feces, staff hands or facility equipment and machinery (Amir et al., 2017; Aslam et al., 2003; Schlegelová et al., 2004).
This study evaluated the antibiotic resistance patterns of Escherichia coli and Staphylococcus aureus isolated from the meat and feces of three commonly hunted/harvested game species from different regions in South Africa.
Methods
All animals were hunted and sampled according to the standard operating procedure approved by the Stellenbosch University Animal Care and Use Committee (ethics number: SU-ACUM14-001SOP).
All three farms are private game farms where the wildlife are free to roam with the other wildlife species on the farm. The wildlife graze and drink on the farm’s natural resources and are given supplementary feed when necessary, especially in times of drought when the grass has become depleted.
Meat (the infraspinatus muscle) and fecal (from the ileum of the small intestine) samples were collected from the same animal in order to directly compare the antibiotic resistance profiles of the meat and feces of each animal. All samples from the same farm were collected on the same day. After sample collection, all samples were stored frozen at –20°C until analysis commenced.
Impala (Aepyceros melampus) meat (n=5) and fecal samples (n=5) were collected from five different impala from a farm in Modimolle, Limpopo, South Africa. Bontebok (Damaliscus pygargus) meat (n=5) and fecal samples (n=5) were collected from five different bontebok from a farm in Wellington, Western Cape, South Africa. Springbok (Antidorcas marsupialis) meat (n=5) and fecal samples (n=5) were collected from five different springbok from a farm in Witsand, Western Cape, South Africa.
Samples were defrosted at room temperature until thawed before analysis commenced. The bacteria were isolated from the samples using a series of plating on selective agar media. The inoculated agar plates were incubated overnight at 35°C. For E. coli, Violet Red Bile Dextrose agar (Merck Bioloab, Modderfontein, South Africa) and then Eosin Methylene Blue agar (Oxoid, Hampshire, UK) was used to isolate the bacteria from the fecal and meat samples. Baird Parker agar (Oxoid, Hampshire, UK) was used to isolate S. aureus from the meat samples. Characteristic growth of E coli on Violet Red Bile Dextrose agar is purple/red colonies surrounded by a halo and characteristic growth on Eosin Methylene Blue agar is colonies with a dark purple center with a green metallic sheen. S. aureus characteristic growth on Baird Parker agar is black colonies with a clear halo.
Gram’s stain and various biochemical tests, the citrate test, catalase test and Staphylase test (Oxoid, Hampshire, UK) were used to confirm colony identity.
After pure cultures were obtained, colonies from the selective agar plates were streaked onto Nutrient agar (Merck Biolab) plates and incubated overnight at 35°C, which were then used for the antibiotic susceptibility test (AST).
Antibiotic susceptibility of all samples were tested in triplicate as each animal was sub-sampled three times in the isolation step. Fresh cultures grown overnight on Nutrient agar (Merck Biolab) were used for the antibiotic susceptibility analysis. The disk diffusion method was used according to the Clinical & Laboratory Standards Institute (CLSI, 2018) 2018 guidelines (M100S) using Mueller-Hinton agar (Merck Biolab). For E. coli, the antibiotic discs (Oxoid, Johannesburg, South Africa) ampicillin 10 μg, chloramphenicol 30 μg, ceftazidime 30 μg, streptomycin 10 μg, sulphafurazole 300 μg and tetracycline 30 μg were tested. For S. aureus, the antibiotic discs (Oxoid, Johannesburg, South Africa) cefoxitin 30 μg, erythromycin 15 μg, oxacillin 1 μg, penicillin 10 U, tetracycline 30 μg and vancomycin 30 μg were tested. E. coli ATCC 25922 and S. aureus ATCC 25923 (Thermo Fisher Scientific, Lake Charles, LA, USA) were used as quality controls and an uninoculated agar plate was used as a negative control. The results of the disc diffusion test classified the isolates as resistant, intermediately resistant or susceptible to the selection of antibiotics, according to the zone diameter specifications as listed by the CLSI 2018 guidelines.
A crude extraction method using lysis buffer and boiling was used to extract DNA from fresh overnight broth cultures of the isolated E. coli meat and fecal samples. The ZymoBiomics DNA kit (Inqaba Biotec, Muckleneuk, South Africa) was used according to the manufacturer’s instructions to extract DNA from fresh overnight broth cultures of the isolated S. aureus meat samples.
Polymerase chain reaction (PCR) was used to detect various antibiotic resistant genes which are commonly associated with phenotypic antibiotic resistance. The antibiotic resistant genes detected in the E. coli isolates were: tetA at 502 bp and tetB at 930 bp (tetracycline resistance), sul1 at 433 bp and sul2 at 721 bp (sulphonamide resistance), blaCMY at 1,000 bp (ampicillin resistance) and aadA at 525 bp (streptomycin resistance). The antibiotic resistant genes detected in the S. aureus isolates were: tetK at 1,515 bp, tetL at 229 bp and tetM at 406 bp (tetracycline resistance), vanA at 732 bp and vanB at 647 bp (vancomycin resistance) and blaZ at 498 bp (penicillin resistance). All reactions were performed in duplicate. The primers and reaction conditions used in this study are described by van den Honert et al. (2020), except the blaZ gene, which is the most common gene encoding production of beta-lactamases to hydrolyse penicillin. The blaZ gene was detected using the following primers and reaction conditions: forward primer sequence: 5’-AAGAGATTTGCCTATGCTTC-3’, reverse primer sequence: 5’-GCTTGACCACTTTTATCAGC-3’; 5 min initial denaturation at 94°C followed by 35 cycles of 94°C for 30s, 55°C for 30s and 72°C for 10 min and a final extension step of 72°C for 10 min (Russi et al., 2015).
Gel electrophoresis was performed using 1.2% agarose gel (Lonza SeaKem, Rockland, ME, USA) stained with EZ-Vision® in-gel solution DNA dye (Amresco, Solon, OH, USA). Gel visualisation was performed using the Bio-Rad Gel Doc XR+ System (Bio-Rad, Hercules, CA, USA) in combination with Image Lab Software V5.2.1.
Statistical analysis was performed on the E. coli isolate results to determine if there were any significant differences between the meat and fecal sample antibiotic resistant levels. The statistical analysis was performed using Statistica 13.2 software (Tibco, Palo Alto, CA, USA). The data was analysed using one-way analysis of variance (ANOVA). The main effect was meat versus fecal sample. Significant results were identified by least significant means (LSM) by using a 95% confidence interval (p≤0.05).
Results and Discussion
Antibiotic resistant bacteria and some of the associated antibiotic resistance genes were found in both the fecal and meat samples of the wildlife species from all three of the farms (Figs. 1 and 2). Table 1 shows the phenotype-genotype correlations of the E. coli and S. aureus isolates. Antibiotic resistant genes were detected in all the isolates which were classified as phenotypically resistant to the various antibiotics.
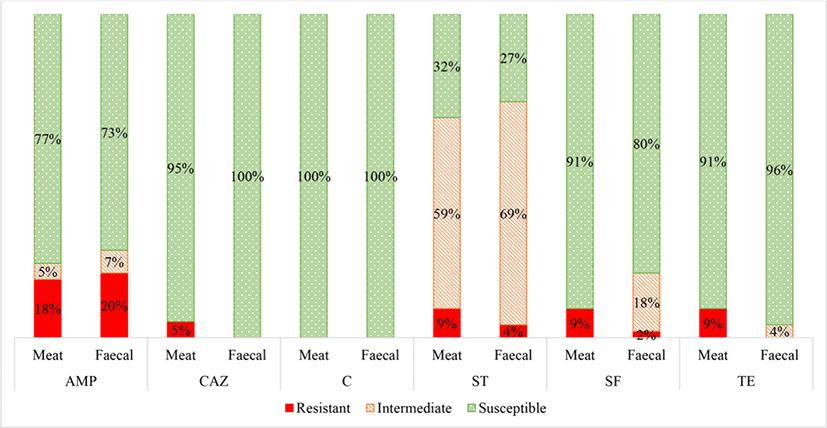
1) AMP, ampicillin (blaCMY gene); SF, sulphonamide (sul1 and sul2 genes); ST, streptomycin (aadA gene); TE, tetracycline (tetA and tetB genes); TE, tetracycline (tetL, tetK and tetM genes); VA, vancomycin (vanA and vanB genes); P, penicillin (blaZ gene); S, susceptible; I, intermediate; R, resistant.
Escherichia coli was isolated from 22 (49%) and 45 (100%) of the meat and fecal samples, respectively. A summary of the AST results for the E. coli isolates is shown in Fig. 1. There were no significant differences in the E. coli antibiotic resistance patterns between the meat and fecal samples, except for ceftazidime. Thus it can be speculated that contamination of the game meat occurred from E. coli both from the carcass feces and from the surrounding environment, equipment and/ or the slaughter personnel (Gouws et al., 2017).
The presence of E. coli (49%) on the meat samples indicates that fecal contamination from the carcass occurred on the meat of these animals during the slaughter process (Aslam et al., 2003). The presence of S. aureus on the meat samples (36%) indicates that unhygienic practices and thus cross-contamination occurred onto the meat from the hide and/ or from the meat handlers. Ultimately, the presence of S. aureus on raw meat is indicative of poor hygiene conditions in the food chain, mainly due to contamination by food handlers and equipment (Naas et al., 2019). The detection of E. coli and S. aureus from the meat samples was expected and the frequency of isolation is similar to those found by Schlegelová et al. (2004), who detected E. coli and S. aureus in meat samples of beef in a slaughterhouse in 67% and 24% of samples, respectively.
The E. coli isolates were most frequently resistant to streptomycin, followed by ampicillin, sulphafurazole, tetracycline and then ceftazidime. No E. coli isolates were resistant to chloramphenicol.
The E. coli isolates were resistant to streptomycin (average 9% meat; 5% fecal) where a high percentage were intermediately resistant (average 59% meat; 68% fecal) (Fig. 1). Other studies have reported that streptomycin resistance is common in food animals due to its extensive use in both agricultural and clinical settings (Boerlin et al., 2005; Bryan et al., 2004; Kozak et al., 2009; Wilkerson et al., 2004). In addition, streptomycin is present in the soil, produced by organisms such as Streptomyces griseus, and could confer a natural low-level resistance to the grazing wildlife (van Overbeek et al., 2002; Wegst-Uhrich et al., 2014).
Moreover, the fecal and meat E. coli isolates were notably resistant to ampicillin (average 18% meat; 20% fecal). More specifically, the high average of ampicillin resistance was mainly attributed to the fecal (100% resistant) and meat (73% resistant) samples from the bontebok species from the Wellington farm (data not shown). The higher ampicillin resistance seen in the bontebok E. coli isolates can be attributed to the fact that this farm which hosts the bontebok was previously a dairy and sheep farm about thirty years ago. The penicillin antibiotic class, which includes ampicillin, is the most widely used antibiotic class in sheep farming (Wegst-Uhrich et al., 2014). The application of antibiotics during the dairy and sheep farming period could have stimulated the development of antibiotic resistant bacteria within the soil which could be transferred to the grazing wildlife (Wegst-Uhrich et al., 2014).
Furthermore, the E. coli isolates had low levels of resistance towards sulphafurazole (average 9% meat; 2% fecal) and tetracycline (average 9% meat; 4% fecal). This is consistent with Li et al. (2007) study which reported similar resistant levels of E. coli isolates from game meat, with sulphafurazole resistance at 7.9% and tetracycline resistance at 13%.
No E. coli meat or fecal isolates were resistant to chloramphenicol. Low resistance was expected as chloramphenicol is not permitted for food animal use in South Africa and many other countries (Rawat and Nair, 2010).
The only significant difference in antibiotic resistance levels between the E. coli isolates from the meat and fecal samples was towards ceftazidime, where 5% of the meat isolates and 0% of the fecal isolates were classified as resistant. Resistance to ceftazidime indicates suspicion for extended-spectrum β-lactamase (ESBL) production (Overdevest et al., 2011). However, additional phenotypic confirmatory tests would still need to be performed to confirm ESBL-production (Dahms et al., 2015; Henton et al., 2011; Overdevest et al., 2011). Other studies have also speculated that environmental ESBL E. coli is a result of human influence, as the majority of ESBLs are reported from human clinical isolates due to the direct use of novel sub-classes of β-lactam antibiotics (Guenther et al., 2011; Skurnik et al., 2006). Thus it can be speculated that contamination of the meat occurred predominantly from human influence, most likely during the skinning and evisceration steps, as also found by Schlegelová et al. (2004), via resistant genotype and phenotype analysis.
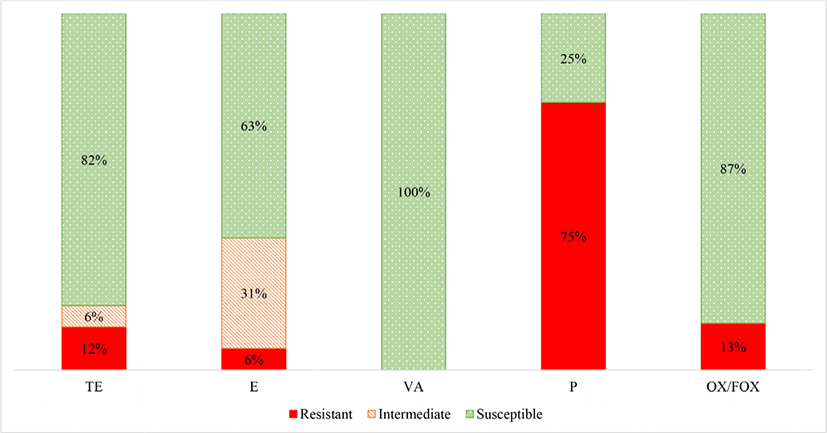
S. aureus was isolated from 16 of meat samples (36%). A summary of the AST results for the S. aureus isolates is shown in Fig. 2. The S. aureus isolates were most frequently resistant to penicillin, followed by oxacillin/ cefoxitin, tetracycline and then erythromycin. No isolates were classified as resistant to vancomycin.
There were 12 S. aureus isolates from the game meat which were resistant to penicillin (75%) (Fig. 2). This was anticipated, as resistance to penicillin is now widespread in humans and animals since the 1960s, in both community, hospital and meat staphylococcal isolates (Appelbaum, 2007; Chambers and DeLeo, 2009; Lowy, 2003; Schlegelová et al., 2004).
Methicillin resistant S. aureus (MRSA) was only detected in two of the bontebok meat isolates (13% average) (indicated by oxacillin and confirmed by cefoxitin). Although genetic confirmation to confirm methicillin resistance should be performed by detection of the mecA gene. Other studies have concluded that contamination of meat with MRSA can result from cross contamination of the carcass from the animal itself or from the people involved in the meat handling during slaughter and processing (Gilmore et al., 2008).
The S. aureus meat isolates were classified as tetracycline resistant in 2 (12%) of the samples. Furthermore, 1 (6%) of the S. aureus isolates from the game meat were classified as resistant to erythromycin, although 5 (31%) were classified as intermediately resistant. Other studies have found varying frequencies of resistance to erythromycin (4.3%–30%) and tetracycline (~50%) of S. aureus from meat (retail non-game meat), where tetracycline resistance is generally more common than erythromycin resistance (Kelman et al., 2011).
None of the S. aureus meat isolates were classified as resistant to vancomycin. Other studies have also reported negligible to very low levels (0%–3%) of vancomycin resistant S. aureus from raw commercial meat samples (Das and Mazumder, 2016; Jackson et al., 2013; Pesavento et al., 2007).
At least one of the selected antibiotic resistant genes were detected in all samples which showed to have a corresponding phenotypic antibiotic resistance pattern. There were seven samples where the antibiotic resistant gene was detected but the phenotypic method classified the isolates as susceptible. This occurred for ampicillin (4) from the E. coli isolates and for tetracycline (2) and vancomycin (1) from the S. aureus isolates. Some possible explanations for these resistance genes being detected in these samples could be that they are inactive genes, meaning that they are present but are not active because there is no antibiotic resistance selective pressure to phenotypically express the gene. Alternatively, PCR can be considered a more sensitive method to the disc diffusion phenotypic method, as resistance is dependent on the size of the zone of inhibition, which is determined by the CLSI committee on an annual basis, whereas resistance in PCR is determined simply by the detection of a resistance gene (Gilmore et al., 2008).
Conclusion
Antibiotic resistant bacteria were detected in the feces and on the raw meat of the wildlife species, with ampicillin and streptomycin resistance being the most prevalent in E. coli from both sample types. The S. aureus isolates from the game meat showed high resistance to penicillin but fairly low resistance to the other five antibiotics.
Although it seems unlikely that antibiotic resistant bacteria would be found in wildlife, movement of antibiotic resistance genes and resistant bacteria can reach these more isolated environments from pollution of human and farm animal environments as well as via supplementary feed and water sources. Contamination via humans during the slaughter and processing steps can also be a source of antibiotic resistant bacteria onto the raw meat.
The E. coli isolated from the meat and feces of the same animal showed to have similar antibiotic resistance patterns except towards ceftazidime, where there was a significant difference in resistance frequencies between the meat and fecal samples. These results indicate that cross contamination of the meat occurred from bacteria both from the carcass and from human origin.
In order to prevent cross-contamination of harmful and/ or antibiotic resistant bacteria from the hides or feces onto raw meat, various precautionary steps can be put in place. For example, exposed muscle must avoid contact with workers hands and the animal’s skin as best as possible. Workers hands are important sources of contamination during processing and thus hand washing is essential in preventing contamination of the carcass. Furthermore, it is important to clean all equipment and meat processing machinery to reduce the effect of cross-contamination.













