Introduction
The study on microbial community in fermented meat products has become a big interest in last decades. Several molecular approaches have been developed to investigate the composition of microbiota in fermented food. The 16S rRNA gene sequencing was frequently used for identifying bacteria in fermented sausages (Blaiotta et al., 2003; Rantsiou et al., 2005; Sanz et al., 1998). Rantsiou and Cocolin (2006) investigated the microbial dynamics in fermented sausage using 16S rRNA gene sequencing during cooking and reported that different genera, species and even strains of bacteria significantly affected its sensory properties. The culture-dependent (agar plating) and culture-independent (direct DNA analysis) combination methods were used to investigate microbial ecology of traditional Italian salami (Silvestri et al., 2007). Lactic acid bacteria (LAB) are the main bacteria found in most fermented meat products. Lactobacillus plantarum, Lactobacillus sakei and Lactobacillus curvatus have been most frequently found species within LAB group. LAB can grow rapidly utilizing carbohydrates in the media where they live and produce lactic acid that lowering pH of the media, even so they also have ability to regulate their cytoplasmic pH to be homeostasis (Cocolin et al., 2001; Hutkins and Nannen, 1993; Leroy et al., 2006). As LAB grow during fermentation, the environment and the composition of indigenous microbiota in traditional dry sausages would be change (Talon et al., 2007b). However, the amount of research investigating the changes in composition of microbiota in fermented sausage made from poultry meat using next-generation of microbial community analysis is still limited.
Fermented meat products such as summer sausage and pepperoni can be found in most parts of the world, even though it is largely produced and consumed in Southern and Central European countries (Campbell-Platt and Cook, 1995). Because meat is one of perishable food, the preservation methods have long been developed. Fermentation and drying have been known as the oldest preservation methods. Fermented sausages are made from cured meat batter, fermented and dried under controlled temperature and relative humidity (RH) conditions in order to extend its shelf life (Leroy et al., 2006; Talon et al., 2007a).
Most fermented sausages are made from pork or beef and mixed with spices. Ground-type meat products are suitable for effective utilization of tough meat obtained from spent layers (Kondaiah and Panda, 1992). However, the availability of fermented poultry based products is still limited in the market concerning the risk of food borne disease caused by Salmonella contamination. To combat the presence of pathogens, sausage can be fermented with selected strain of lactic acid producing bacteria, cooked and dried. These bacteria play role not only to preserve the sausage by increasing its acidity level but also to enhance flavor, aroma and texture through enzymatic processes (Caplice and Fitzgerald, 1999; Urso et al., 2006). Furthermore, the use of natural ingredients that could inhibit the growth of pathogens is required. Chopi (Zanthoxylum piperitum) is one of spices that is available in East Asia and used traditionally as herbal medicine. The essential oil and methanolic extract of chopi have been confirmed having antimicrobial and antioxidant activity against some food pathogens and lipid oxidation in meat homogenate (Choi et al., 2008; Park et al., 2008; Tanabe et al., 2002; Yamazaki et al., 2007). Therefore, the objective of this study was to observe the effect of ground chopi addition on the changes in physicochemical traits and composition of microbiota during different manufacturing stages based on their molecular profiles, using high-throughput sequencing on Illumina MiSeq.
Materials and Methods
Frozen deboned skinless thighs from 70-week-old spent layer (Hy-Line) were thawed overnight at 2±2°C and cold ground twice through 10-mm plate using a meat grinder (PC-6 5-F, Sung Chang Co., Ltd., Korea). Thigh meat was chosen considering its high myoglobin content compared with breast meat and the addition level (%) of other ingredients was based on weight of the ingredient to weight of ground meat (w/w). Ground thigh meat was mixed for 5 min using a HeavyDuty 5KPM50 mixer (KitchenAid, USA) with salt (2.3% w/w), sodium erythorbate (0.05% w/w), dextrose (0.75% w/w), sucrose (1% w/w), ground black pepper (0.7% w/w), all spices (0.77% w/w), carrageenan (1.0% w/w), hickory liquid smoke (0.1% w/w), and curing salt (0.3% w/w) consisting of sodium chloride (93.1%), sodium nitrite (5.9%) and sodium carbonate (1.0%). Freeze-dried starter culture (0.2% w/w) consisted of Staphylococcus carnosus, Staphylococcus xylosus, Lactobacillus curvatus and Lactobacillus sakei (Lyocarni VBL-97, Sacco System, Italy) was rehydrated in distilled water (1:1 w/v) prior to mix with the batter. Ground dried-chopi fruit (Zanthoxylum piperitum) purchased from local market was added at different levels; 0% (control), 0.1%, 0.3% and 0.5% (w/w) and mixed for another 5 min. The sausage batter was stuffed (20-cm long/sausage link) into 60-mm (Ø) cellulose casing (Teepak Co., USA) and was hanged and processed in a chamber (VC 2057, Votsch Industruetechnik, Germany) with controlled temperature and relative humidity (RH). Sausages were fermented until reaching pH of 5.0 at 35°C/RH 73% for 15 h, stepwise cooked at 54°C/RH 27% for 1 h, 63°C/RH 31% for 1.5 h, 74°C/RH 17% for 1.5 h, 77°C/RH 15% for 3 h, 77°C/RH 31% for 5 min and cooled down at 10°C/RH 80% for 1 h totaling 8 hours for this stage. The cooked sausages were dried for 7 d. The 7-d of drying process was started at 10°C/RH 80% for 48 h, 10°C/RH 75% for 48 h and 10°C/RH 70% for 72 h. Sampling (n=3) for physicochemical and microbiological analyses was performed at the beginning of the manufacture (raw batter), the end of fermentation, cooking (after cooling down) and drying.
The sample (5 g) and distilled water (45 mL) were homogenized (PH91, SMT Co., Ltd., Japan) at 10,000 rpm for 1 min and the mixture was used for pH determination. Changes in pH were recorded in triplicate on each manufacture stage (raw, fermentation, maturation and drying) using a pH meter (SevenEasy pH, Mettler-Toledo GmbH, Switzerland) calibrated with acid (pH 4.01), neutral (pH 7.00) and alkali (pH 9.21) technical buffer solutions (Seven Easy pH, Mettler-Toledo GmbH, Switzerland) at 22°C according to manufacturer’s instruction. Water activity was measured with aquaspector (AQS-2, Nagy, Germany) in triplicate for each 5 g of samples.
The instrumental color was recorded in five repetitions by measuring International Commission on Illumination’s system for lightness (CIE L*), redness (CIE a*) and yellowness (CIE b*) using a chromameter (CR-400, Konica Minolta Inc., Japan). The light source of illuminant C (2° observer) with 8 mm aperture and attached-closed cone was calibrated using a white plate (Y=93.6, X=0.3134, y=0.3194).
Lipid oxidation was analyzed using the 2-thiobarbituric acid reactive substances (TBARS) method by Sinnhuber and Yu (1977) with slight modification. A total of 0.5 g of sample was prepared in triplicate and vortex-mixed with 0.1 g of antioxidant mixture (consisting of 54% propylene glycol, 40% Tween 20, 3% butylated hydroxytoluene, and 3% butylated hydroxyanisole), 3 mL of 1% TBA (thiobarbituric acid) in 0.3% NaOH, and 17 mL of 2.5% trichloroacetic acid in 36 mM HCl (Sigma-Aldrich Corp., LLC., USA). The sample was heated in a water bath (BW-20G, Biotechnical Services, Inc., USA) at 100°C for 30 min and then immersed in ice water for 15 min. Subsequently, 5 mL of aqueous sample was mixed with 3 mL of chloroform (Daejung Chemical and Metals Co. Ltd., Korea) to remove the dirt. The absorbance value of the upper layer was recorded at 532 nm (UV Mini 1240 PC, Shimadzu Corp., Japan) against blank after centrifuging at 2,400 g for 30 min at 4°C (1248R, Labogene, Denmark). The result was expressed in mg of malondialdehyde (MDA) per kg of sample.
For bacterial counts, 1 g of sample was put in a sterile bag and prepared in triplicate (Whirl-Pak, Nasco, USA). Sample was homogenized with 9 mL of sterilized 0.1% peptone in distilled water for 2 min using a stomacher (400, Seward Ltd., UK). Decimal dilutions were prepared using sterilized 0.1% peptone water. Total viable counts (TVC), LAB and coliform were enumerated using plate count agar, De Man, Rogosa and Sharpe agar and violet red bile agar, respectively (Difco, Becton, Dickinson and Company, USA). The LAB plates were incubated in anaerobic chambers with anaerobic atmosphere generation bags (Sigma-Aldrich Corp., LLC., USA). For TVC, the plates were incubated at 37°C for 48 h. While, both coliform and LAB plates were incubated at 35°C for 24 to 48 h. Microbial population was counted and expressed as log CFU/g.
Sausage samples were processed for DNA extraction. NucleoSpin Soil kit (MN, Düren, Germany) was used to extract genomic DNA of microorganisms in the samples. PCR was conducted to amplify 16S rRNA gene using barcoded primers targeting V4 region. The primers have the following sequences, respectively: 5'-GGACTACHVGGGTWTCTAAT-3 and 5'-GGACTACHVGGGTWTCTAAT-3'. PCR reaction solution contained genomic DNA (5 ng), reaction buffer (25 mM Mg2+), dNTP (200 μM each), Ex taq (0.75 unit, Takara, South Korea), and barcoded primers (5 pmole each). PCR reaction was carried out as follows; 94°C for 3 min for the initial denaturation, 30 cycles of 45 s at 94°C, 1 min at 55°C and 90 s at 72°C for amplification, and 72°C for 10 min for the final extension. The same amount of each PCR amplicon was pooled, and the pooled amplicons were subjected to library construction for Illumina MiSeq sequencing using NEBNext® UltraTM DNA Library Prep Kit (NEB, Cat. E7370S). The library was sequenced for paired-end 250-bp reads in the Illumina MiSeq.
Raw MiSeq reads were processed to be demultiplexed and trimmed by in-house perl scripts. The processed paired reads were concatenated, and the concatenated reads were applied to quantitative insights into microbial ecology (QIIME) for microbial analysis version 1.9.1 (Caporaso et al., 2010). Greengenes database (gg_ptus-13_8-release version, 97% nucleotide identity) was used to assign each concatenated read to operational taxonomic unit (OTU) (DeSantis et al., 2006). The α- and β-diversity, the cluster pattern and summarized taxonomy of each sample were analyzed using the QIIME pipeline after OTU assignment. Principal coordinate analysis (PCoA) was performed based on unweighted and weighted UniFrac distances. The data of physicochemical properties and bacterial counts were subjected to two-way analysis of variance (ANOVA). Chopi addition level (control, 0.1%, 0.3%, and 0.5%) and manufacture stage (raw, fermentation, cooking, drying) were used as factors. Interaction effects were found in observed variables except water activity (only different manufacture stage effect was found). Therefore, the means of treatment groups were then separated using Duncan’s multiple range test (p<0.05) according to chopi level and manufacture stage with SAS 9.1 (SAS Institute Inc., USA).
Results and Discussion
The declines in pH and water activity of the chicken summer sausage during manufacture are shown in Table 1. The effect of ground chopi addition on pH was observed after cooking and drying stages. However, the addition of ground chopi did not influence water activity of the sausage during manufacture. Samples added with ground chopi had lower pH than control after cooking and drying stages, while no differences were found at the beginning of the manufacture and after fermentation. The initial change of pH started during fermentation. The pH dropped about 1.50 to 1.56 unit, reached the lowest value (4.83–4.91) after cooking, and no significant changes were observed until 7 d of drying. Drying for 7 d lowered not only the pH but also water activity of the sausage significantly. The water activity values showed slow decreases. The final product obtained after drying had the lowest water activity of 0.91, regardless of chopi addition level.
Changes in pH and water activity are two basic elements in fermented sausage manufacturing. The accumulation of lactic acid from carbohydrate metabolism by lactic acid bacteria during fermentation and the length of drying time determine the decline trend of pH value and water activity. Deumier and Collignan (2003) reported that the final pH of dry chicken summer sausage was about 4.97, while the water activity of 0.82 was recorded on day 24 of drying. These mean that our samples should be dried longer to achieve water activity below 0.90.
The instrumental color of the sausage changed during manufacture and the addition of ground chopi at 0.3% and 0.5% slowed down (p<0.05) the redness development (Table 1). Fermentation process increased CIE L* value (p<0.05) in all experimental groups and the value decreased signifiantly after cooking. The CIE a* value of all groups was significantly higher than that of raw samples after fermentation. The final products of the control and 0.1% chopi group had significantly higher CIE a* value than those of 0.3% and 0.5% group. The addition of ground chopi into raw sausage batter reduced yellowness (p<0.05) as it also reduced the lightness (p<0.05). However, no significant differences were found on yellowness between control and added chopi groups after fermentation and so forth because the CIE b* value of added chopi groups increased (p<0.05) and being close to that of control. The dark appearance of the ground dried-chopi might reduce the lightness and yellowness of the raw sausage batter. Further, the development of redness during manufacture and the manufacture process itself could influence the lightness and yellowness of the final products.
The development of redness in fermented cured meat products is merely caused by the use of nitrate/nitrite salts. Under acid condition, the microorganism activates nitrate/nitrite reductase resulting in the generation of nitric oxide (NO). NOs bind to the heme pigment of myoglobin and the NO-heme pigment complexes result in the red appearance of the cured meat (Pegg and Honikel, 2014). In this study, lower CIE a* value found in the sausages added with ground chopi during manufacture could be caused by the antimicrobial activity of the ground chopi itself. The essential oil of Zanthoxylum piperitum possesses inhibitory activity against food borne pathogens and oral Streptococcus mutans (Choi et al., 2008; Park et al., 2008). Therefore, the number of nitrate/nitrite reductase-positive bacteria might be affected by ground chopi and less NO was produced than control.
Changes in TBARS value of the sausage during manufacture were also affected by the addition of ground chopi (Table 1). TBARS value increased (p<0.05) during manufacture in all groups. However, the final products containing ground chopi had lower TBARS value than that of control. Yamazaki et al. (2007) reported that the methanolic extract of Zanthoxylum piperitum fruit had an antioxidant activity equal to that of α-tocopherol with hyperoside (quercitin-3-O-galactoside) and quercitrin (quercitin-3-O-rhamnoside) as the main compounds. Moreover, hyperoside and quercitrin prevented cell damage induced by oxidation in vitro and in vivo (Babujanarthanam et al., 2011; Piao et al., 2008). The antioxidative effect of Zanthoxylum piperitum pepper in pork homogenate against lipid oxidation has also been proved in previous study (Tanabe et al., 2002). Present study confirms that the addition of ground chopi slow down lipid oxidation in chicken summer sausage during manufacture. Among chopi groups, addition level of 0.3% and 0.5% (w/w) resulted in lower TBARS value in the final product than the others.
Total viable count, lactic acid bacteria and coliform population were influenced by different manufacture stages and addition level of ground chopi (Table 2). Total viable count, lactic acid bacteria and coliform population reduced significantly at the end of manufacture. However, only lactic acid bacteria grew (p<0.05) during fermentation, thus the population of lactic acid bacteria outnumbered total viable count (aerobic bacteria) at the end of fermentation. Even so, the population of lactic acid bacteria started declining after cooking and further until the end of drying. The addition of ground chopi significantly reduced total viable count after fermentation and further until the end of drying. The population of lactic acid bacteria was only affected at the beginning of manufacture, fermentation and cooking, while no significant differences were found after drying. Similar declining trend was also found in the population of coliform as affected by ground chopi. The addition of ground chopi at 0.5% reduced (p<0.05) the population of coliform more than lower addition levels after cooking. At the end of drying, no differences were found between the addition level of 0.3% and 0.5% on total coliform. Park et al. (2008) reported that the essential oil extracted from chopi exhibited antimicrobial activity against Bacillus cereus, Staphylococcus aureus, Salmonella cholerawsuis, and Vibrio parahaemolyticus with limonene as the major compound. The antimicrobial activity of limonene has been confirmed in model bacteria membranes through membrane fluidization (Hąc-Wydro et al., 2017).
The reduction of α-diversity (species richness) of the chicken summer sausage during manufacture is shown in Fig. 1. The rarefaction curves (Fig. 1A) peaked at 5,000 sequences per sample highlighting that an adequate sequencing depth and observed species (OTUs) discovery was achieved or suggesting that the majority of bacterial phylotypes present in samples were successfully identified. Significant differences in averaged OTUs from 0 to 10,000 sequencing reads were found between raw and further stages (Fig. 1B). However, no differences were found after fermentation and further until the end of manufacture. The significant effect of ground chopi was found at the beginning of manufacture and during fermentation. The addition of ground chopi reduced species richness in raw and fermented stages.
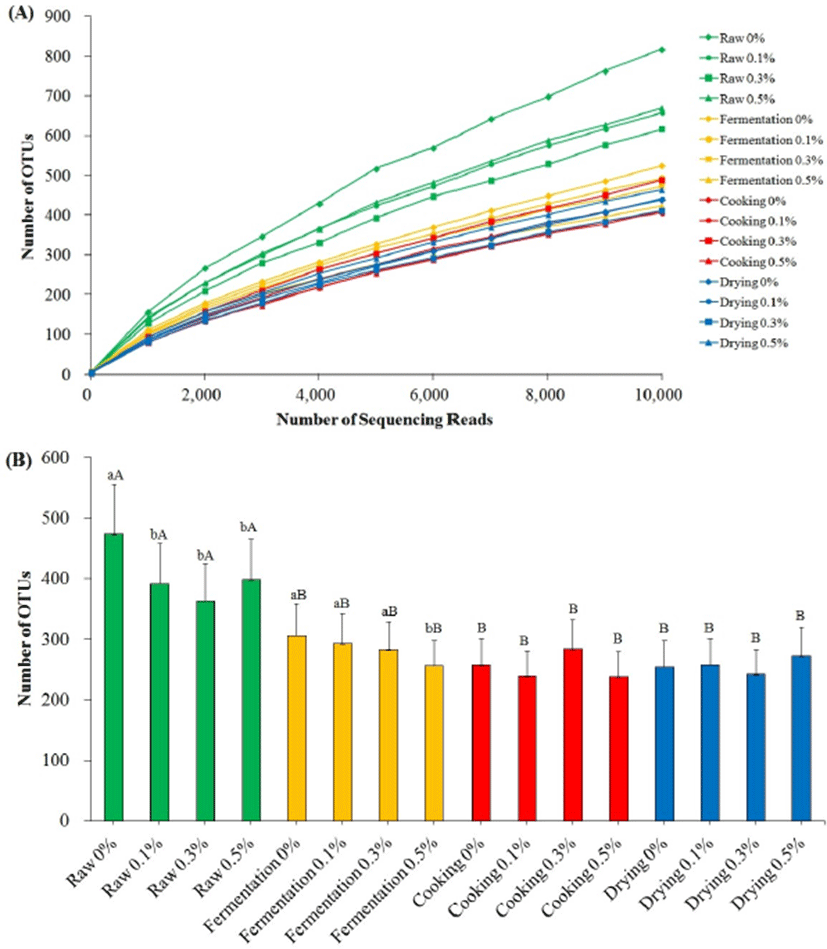
The changes in environmental conditions of the sausage during manufacturing led the decreases of species richness. Anaerobic and low pH conditions in fermentation stage were fit only for some groups of bacteria to grow such as lactic acid-producing bacteria. These bacteria are predominant in most of fermented food because of their ability to live in low level of oxygen environment, metabolize carbohydrates to be lactic acid, maintain their inner cell pH and produce a biological weapon against their opponent called bacteriocin. The other bacteria might be not suitable with those conditions (Cocolin et al., 2001; Hutkins and Nannen, 1993; Leroy et al., 2006). Therefore, fermentation lowered the species richness of fermented sausage. The pH of the sausage reflected the species richness and predicted the composition of microorganism during manufacture. The species richness is dependent on the level of pH due to the excessive growth of lactic acid bacteria. These findings are in accordance with Fontana et al. (2005), Park et al. (2012) and Roh et al. (2010). Moreover, this study confirms the antibacterial effect of ground chopi in chicken summer sausage during fermentation process, however, with unclear effect further until the end of drying process because cooking might also diminish the presence some bacteria.
In this study, culture-independent method was used to analyze the microbial community profiles of fermented sausage made from spent layer meat. Classic agar plating, PCR and denaturing gradient gel electrophoresis were frequently used for microbial analysis of fermented sausages in previous studies with 93%–100% sequence similarity cut-off value (Aquilanti et al., 2007; Cocolin et al., 2001; Janssens et al., 2012; Silvestri et al., 2007). However, DNA sequencing on Illumina MiSeq costs cheaper with high-throughput results (Caporaso et al., 2012).
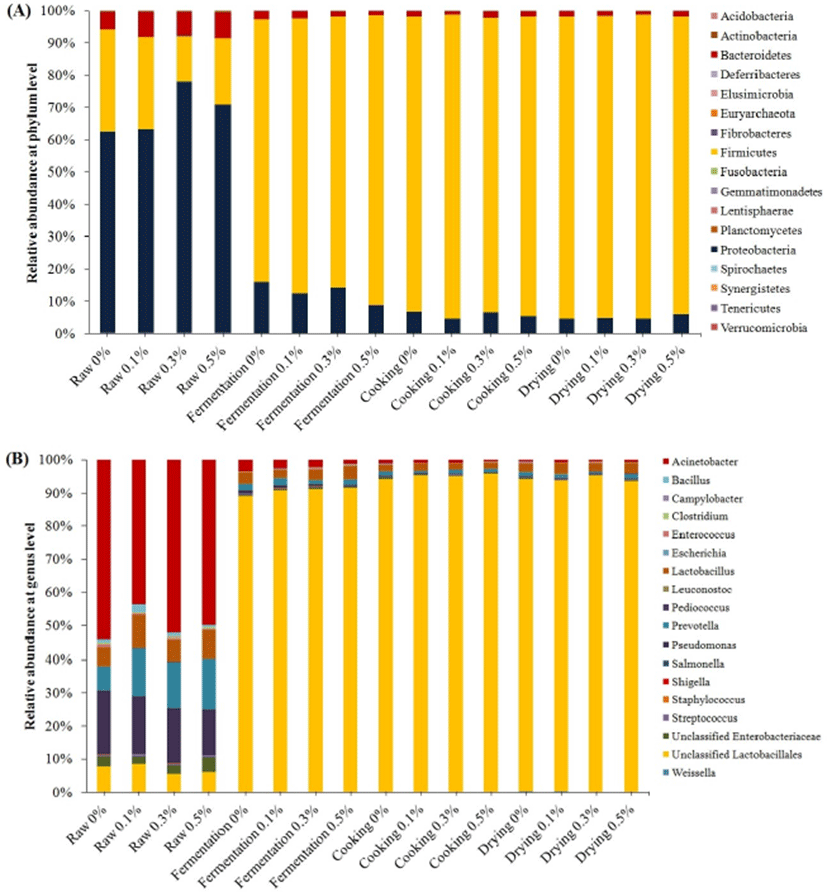
The 16s rRNA gene sequencing was used to determine the phylogenetic classification of microbial community in chicken summer sausage taken from each manufacturing stage. The changes in composition of microbiota at phylum and genus level during manufacture are shown in Figs. 2A and B, respectively. Although ground chopi significantly reduced the number of total viable count and coliform after fermentation, the effect of ground chopi addition was not clear on the proportion of microbiota as it was also not clear on species richness. The three major (>1%) phylum were identified; Bacteroidetes, Firmicutes and Proteobacteria, using QIIME pipeline. In raw samples, Proteobacteria was dominant among other major phylum with proportion more than 60%, followed by Firmicutes (20%–25%) and Bacteriodetes (<10%). However, only Firmicutes increased significantly after fermentation and peaked at drying stage, dominating almost 90% of total identified microbiota at both cooking and drying stages. After fermentation and so forth, Bacteriodetes and Proteobacteria population decreased sharply. At genus level, lactic acid bacteria was the predominant group of bacteria after fermentation and so forth, highlighted by a significant escalation of unclassified Lactobacillales and Lactobacillus. The proportion of other lactic acid bacteria such as Enterococcus, Leuconostoc, Lactococcus, Pediococcus, Streptococcus and Weisella did not change significantly during manufacture. The proportion of Acinetobacter, Prevotella, Pseudomonas and Unclassified Enterobactericeae reduced significantly after fermentation and so forth. Although Staphylococcus was not a predominant genus found during manufacture, its proportion increased during fermentation. However, cooking and drying lowered the proportion of Staphylococcus significantly. Janssens et al. (2013) reported that coagulase-negative staphylococci, for instance Staphylococcus xylosus, Staphylococcus carnosus, Staphylococcus saprophyticus and Staphylococcus equorum, were naturally present in the raw meat batter of fermented sausages or added from the starter culture. These bacteria play role in flavor enhancement of fermented sausages and inhibit oxidation. However, they are sensitive to long exposure of acidic environment.
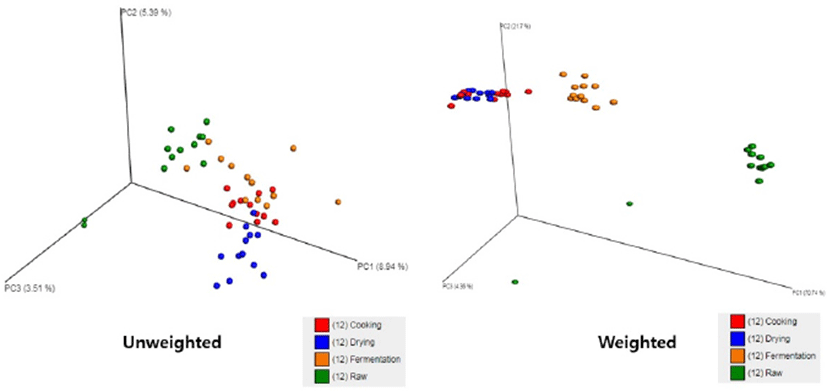
The β-diversity of microbial profiles among each stage of manufacture was analyzed using QIIME pipeline based on the distance of population profiles. Population profile distance of each stage is shown in Fig. 3. The closest relationship was observed between cooking and drying group, while the distance between raw-fermentation, raw-cooking, raw-drying, fermentation-cooking and fermentation-drying were far from each other. Environmental changes during manufacturing were the main factor of these differences. Temperature and relative humidity of the chamber were similar for both maturation and drying stages. Therefore, the profiles of microbiota obtained from those stages were almost similar. Raw samples consisted of all kind of bacteria that naturally found in chicken meat and other natural ingredients. However, fermentation was suitable only for some species of bacteria to grow. Lozupone et al. (2007) mentioned that the microbial community in different environmental conditions varied according to quantitative and qualitative β-diversity analysis.
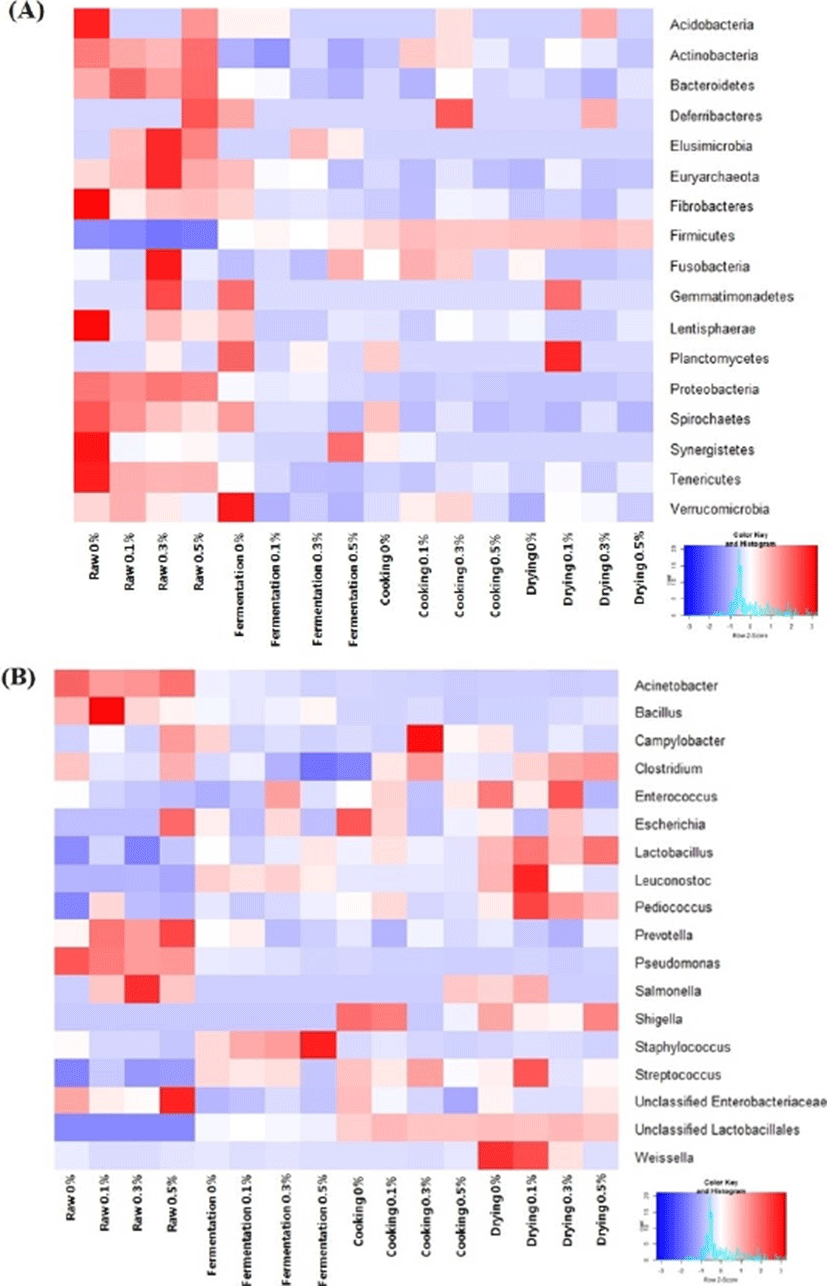
The clustering patterns were figured out through heatmap analysis at phylum (Fig. 4A) and genus level (Fig. 4B). The proportions of microorganism at phylum and genus level that increased or decreased during manufacture were subjected into heatmap analysis. The population of anaerobic gram-positive such as Lactobacillus, Lactococcus, Leuconostoc, and other unclassified Lactobacillales increased after fermentation. In contrast, pathogen and spoilage bacteria such as Enterobacter, Prevotella, and Pseudomonas decreased after fermentation. The less abundant bacteria with low population such as Acenitobacter, Bacillus, Campylobacter, Clostridium, Escherichia, Enterobacter, Pseudomonas, Samonella, Shigella and the absence of Listeria in the final product highlighted that the manufacturing met a high hygienic quality standard even though the addition of ground chopi could not diminish the presence of Clostridium and Shigella at the end of manufacture. These bacteria have been known to occur and most frequently isolated from raw meat (Nychas et al., 2008). The proliferation of lactic acid bacteria contributes to inhibit and control the growth of some pathogenic bacteria by producing acids and bacteriocins in meat and meat products (Castellano et al., 2004; Kouakou et al., 2009). For instance, plantaricin produced by Lactobacillus plantarum can prevent food spoilage and inhibit pathogenic bacteria (Leal-Sanchez et al., 2002). Talon et al. (2007b) mentioned that formula variation, fermentation and cooking conditions were related to the composition of microbiota in dry fermented sausage and lactic acid bacteria constituted the major microbiota at the end of cooking stage. However, the information regarding characterization at species level of those lactic acid bacteria was not performed in present study.
Conclusion
Ground chopi (Zanthoxylum piperitum) delayed lipid oxidation and reduced the number of spoilage bacteria in chicken summer sausage during manufacture with unclear effect on microbial diversity after cooking stage. Fermentation lowered the species richness and diversity, which were dynamically responded to the abundance of lactic acid bacteria. The microbial community differed among different stages of fermented sausage manufacture according to β-diversity analysis and raw sausage batter was observed having the largest diversity according to α-diversity analysis. These findings have been showing implications for understanding the relationship between microbial community in fermented sausage and the environmental factors, such as ingredients (ground chopi addition), temperature and pH during manufacture. In addition, the high-throughput sequencing using Illumina MiSeq and microbial community analysis using QIIME pipeline could identify the changes in composition of microbiota in summer sausage made from spent layer meat at all stages of manufacture. While additional studies will be needed to fully analyze the microorganisms at species level.













