Introduction
Escherichia coli (E. coli) O157:H7 is a significant foodborne pathogen that causes bloody diarrhea and occasionally, hemolytic-uremic syndrome (HUS) (Abuladze et al., 2008). These illnesses have been related to ground beef, fruit, vegetables, and undercooked food (Ackers et al., 1998). On December 24, 2009, the United States Department of Agriculture Food Safety and Inspection Service (FSIS) issued a recall of 248,000 pounds of beef products from steak and poultry that have been contaminated with E. coli O157:H7 in Colorado, Iowa, Kansas, Michigan, South Dakota, and Washington (CDC, 2010).
Pathogenic E. coli caused 291 foodborne illness outbreaks from 2011 to 2012, accounting for 24.1% of total food poisoning outbreaks in Korea (Oh et al., 2013). According to the statistics from the Korea Centers for Disease Control and Prevention, foodborne outbreaks of pathogenic E. coli were frequently associated with the contamination of pickled vegetables, beef, pork, and drinking water in summer (Oh et al., 2013). In 2012, kimchi contaminated by E. coli O169 caused serious cases of enteritis in schoolchildren (Cho et al., 2014). Although enterotoxigenic E. coli is more prevalent than enterohemorrhagic E. coli, there is a significant risk for the contamination of E. coli O157:H7 in meat from livestock animals (Cho et al., 2014; Kang et al., 2004).
Bacteriophages are viruses causing lysis of the host bacteria. Since its discovery by Twort in 1915 and d’Herelle in 1917, bacteriophage has been exploited to control bacteria (Clokie and Kropinski, 2009; Kutter and Sulakvelidze, 2004). With the emerging public health concern about antibiotic resistant bacteria, the use of bacteriophages to control pathogenic bacteria has received more attention by the food industry and medical science (Clark and March, 2006; Sillankorva et al., 2012; Verraes et al., 2013). The U.S. Food and Drug Administration (FDA) approved Listex P-100 (Micros Food Safety, Netherlands) and EcoShield (Intralytix, Inc., USA) for commercial use to clean hard surface in food processing plant and to reduce the risk of meat contamination by Listeria monocytogenes (L. monocytogenes) or E. coli O157:H7, respectively. In addition, previous studies showed the inhibitory effect of bacteriophage cocktails on E. coli O157:H7 on hard surfaces and in tomato, spinach, ground beef, and meat (Abuladze et al., 2008; O'Flynn et al., 2004).
It was demonstrated that E. coli O157:H7 was reduced in cooked and raw beef by bacteriophage treatment (Hudson et al., 2013; Hudson et al., 2015). However, bacteriophages that target E. coli O157:H7 were not used in pork and chicken meat. Therefore, the aims of this study were to determine the optimal treatment of bacteriophage to control E. coli O157:H7 and to investigate the inhibition of E. coli O157:H7 inoculated on beef, pork, and chicken by bacteriophage.
Materials and Methods
E. coli O157:H7 ATCC 43888, E. coli O157:H7 ATCC 43889, and E. coli O157:H7 ATCC 43890 were purchased from American Type Culture Collection (ATCC; USA). Host bacteria were grown in Luria Bertani (LB, Sigma-Aldrich, USA) broth on shaking incubator (SI-600R, Jeiotech, Korea) at 37℃ overnight.
Fourteen lytic bacteriophages against E. coli O157:H7 ATCC 43889 were obtained from the bacteriophage bank of Hankuk University of Foreign Studies in Korea (Table 1). The titration of each bacteriophage strain was determined by the plaque assay, as previously described (Hudson et al., 2013; Hudson et al., 2015). Briefly, 100 μL of E. coli O157:H7 and 100 μL of bacteriophage were diluted 10-fold with SM buffer (0.05 M Tris-HCl, 0.1 M NaCl, 0.008 M MgSO4, 0.01% gelatin, pH 7.5), mixed in a 1:1 ratio and incubated for 1 h in a shaking incubator (SI-600R, Jeiotech, Korea) at 150 rpm. Subsequently, the mixtures were inoculated in LBC soft agar (LB broth, 0.1% CaCl2, 0.6% agar). LBC soft agar was overlaid onto a substrate of LBC agar (LB agar, 0.1% CaCl2) and incubated at 37℃ for 18 h. Five milliliter of SM buffer were added to each plate exhibiting a sufficient number of bacteriophage plaques and plates were transferred to an orbital shaker (OS-752, Optima, Japan) at 100 rpm for 3 h. The supernatant was collected with a syringe (KOVAXSYRINGE 5 mL, Korea vaccine, Korea) with intermittent agitation and filtered through a 0.2 μm filter (minisart CA 16534, Sartorius Stedim Biotech A.G., Germany). For bacteriophage propagation, 500 μL of E. coli O157:H7 were inoculated into 100 mL LBC broth with 1.5 mL of 30% glucose (Sigma-Aldrich) and incubated at 37℃ for 1 h. Subsequently, bacteriophages were inoculated into the E. coli/LBC broth mixture, incubated at 37℃ for 5 h and the cultured broth filtered through a 0.2-μm syringe filter. The final titer of bacteriophages was adjusted to 109-1011 plaque forming unit (PFU)/mL and bacteriophages stored at 4℃ until further use.
For comparing the lytic activity of each bacteriophage strains, E. coli O157:H7 was treated with 10,000 multiplicity of infection (MOI) used in previous studies (Hudson et al., 2013; Hudson et al., 2015). Bacteriophage activity was measured by challenge test, as described, with slight modifications (O'Flynn et al., 2004). Briefly, E. coli O157:H7 was cultured in LB broth at 37℃ overnight and diluted to 1×105 colony-forming unit (CFU)/mL with 0.1% peptone water. The titer of each bacteriophage strain was adjusted to 1×109 PFU/mL with SM buffer. Each bacteriophage strain was mixed with E. coli O157:H7 and transferred to a shaking incubator at 37℃. After 10, 20, 40, and 60 min incubation, 10-fold diluted samples were plated on SMAC and incubated at 37℃ overnight and the populations of E. coli O157:H7 were counted.
Based on the comparison of lytic activity between bacteriophage strains, BPECO19 showed the strongest lytic activity against E. coli O157:H7 in this study. To examine the MOI-dependent inhibition of BPECO19, 1×105 CFU/mL of E. coli O157:H7 was treated in vitro with 10, 100, 1,000, and 10,000 MOI of bacteriophage BPECO19 at 37℃ for 0, 10, 20, 40, and 60 min. Subsequently, each sample was 10-fold diluted and plated. After overnight incubation at 37℃, the population of host bacteria was counted.
Beef, pork, and chicken meat were purchased at a local market (Korea). Each meat sample was aseptically cut to squares of 2 × 2 cm. Experimental design and treatment time were modified from previous studies (Abuladze et al., 2008; Bigwood et al., 2008; Hudson et al., 2013; Hudson et al., 2015). Cocktail of E. coli O157:H7 ATCC 43888, E. coli O157:H7 ATCC 43889, and E. coli O157: H7 ATCC 43890 at a concentration of 1×105 CFU/cm2 were inoculated on the surface of each meat sample with pipet spreading. Subsequently, each meat sample was inoculated with 1,000, 10,000 and 100,000 MOI bacteriophage BPECO 19 and stored at 4℃ and 37℃ for 1, 2, 4, 8, 12, 24, 48, 72, 120, and 168 h post-inoculation. Positive control was contaminated with E. coli O157:H7 in meat without bacteriophage treatment. Negative control was not contaminated with E. coli O157:H7 in meat treated with same amount of LB media instead of bacteriophage.
Each experimental group had 3 technical replicates. Three times experiments were performed independently. Data from each group were expressed as mean and standard deviation. Statistical significance was analyzed at p<0.05 by one-way analysis of variance (ANOVA) and paired t-test using Statistical Package for Social Science (SPSS) software v18 (IBM, USA).
Results
The origin and plague size of 14 bacteriophage strains were shown in Table 1. While 12 bacteriophages showed the pin-pointed plague with less than 1-2 mm diameter, 6-8 mm plague of BPECO19 and BPECO 24 was prominently observed (Table 1). When the lytic activity of 14 bacteriophage strains was assessed at 10,000 MOI against E. coli O157:H7 cultured in vitro, 10 bacteriophage strains caused less than 1-2 log reduction/ml at 1 h (data not shown). Among them, bacteriophage ECO3, BPECO6, BPECO18, and BPECO19 significantly reduced E. coli O157:H7 within 1 h (Fig. 1). Especially, bacteriophage BPECO19 decreased E. coli O157:H7 by 4.53 log in 10 min (p<0.05) and E. coli O157:H7 was not detected in 20 min treatment (p<0.001) (Fig. 1). Whereas bacteriophage ECO3 and BPECO18 gradually decreased host bacteria until 60 min (p<0.05) (Fig. 1), bacteriophage BPECO6 reduced E. coli O157:H7 by 2.18 log at 40 min (p<0.05) but the effect was less pronounced after 60 min (Fig. 1). When the plague size and lytic effect of each bacteriophage strain was considered, bacteriophage BPECO19 was selected to control E. coli O157:H7 in artificially contaminated meat samples.
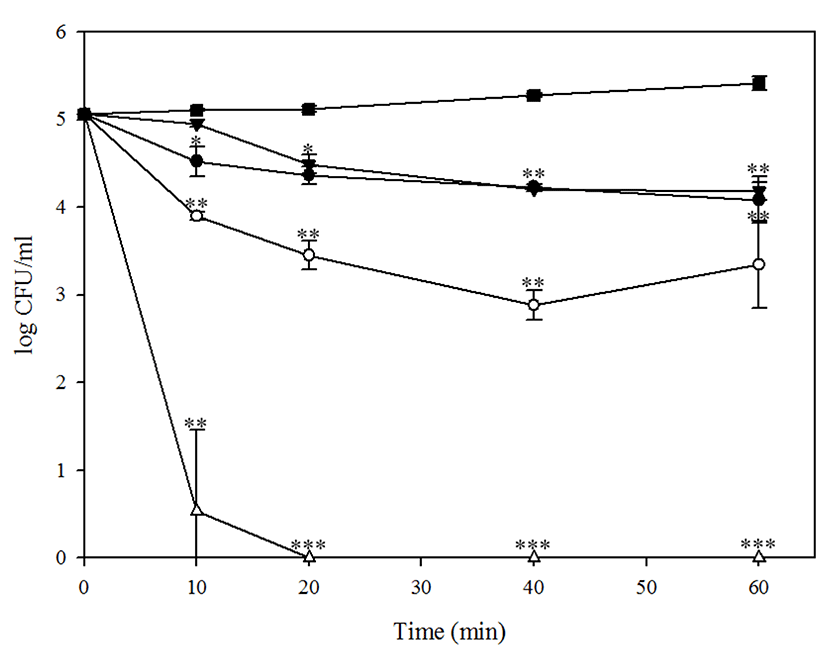
When E. coli O157:H7 was treated in vitro with 10, 100, 1,000, and 10,000 MOI of BPECO19 up to 1 h, MOI-dependent reduction of E. coli was clearly shown in Fig. 2. BPECO19 at 10,000 MOI completely inhibited E. coli O157:H7 in 10 min (p<0.05). BPECO19 at 1,000 MOI gradually reduced E. coli O157:H7 from 5.10 Log CFU/mL to 0.99 Log CFU/mL in 60 min (p<0.05). BPECO19 at 100 MOI decreased E. coli O157:H7 by 0.36 log in 60 min (p<0.05). Although 10 MOI transiently decreased E. coli O157:H7 by 0.29 log in 20 min, host bacteria was not significantly reduced in 60 min (Fig. 2). From this data, treatment condition of bacteriophage BPECO19 to control E. coli O157:H7 in meat sample was determined.
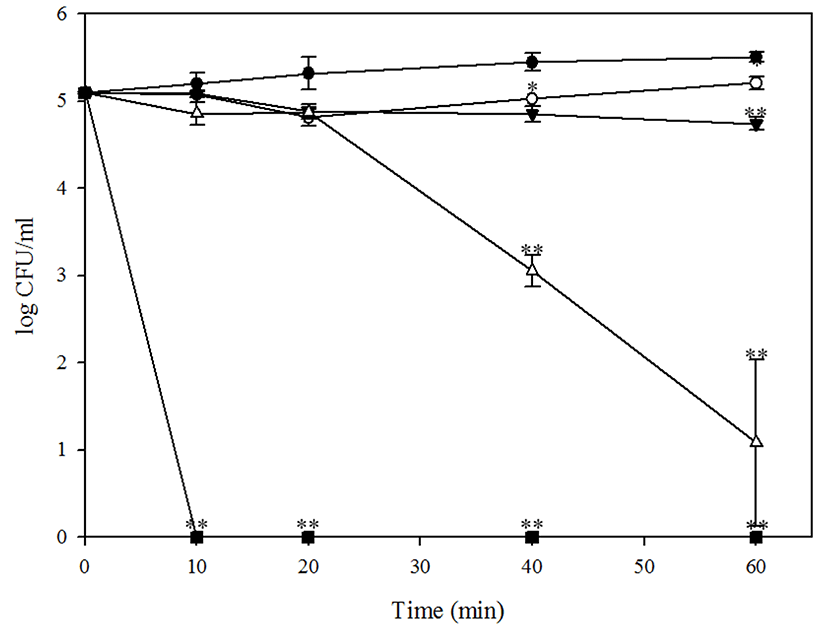
When bacteriophage BPECO19 was applied to artificially contaminated beef at 4℃, E. coli O157:H7 reduced at 1,000, 10,000, 100,000 MOI proportionately. However, BPECO19 could not inhibit E. coli O157:H7 in beef stored at 37℃ (data not shown). E. coli O157:H7 was gradually reduced from 5.09 Log CFU/cm2 to 1.37 Log CFU/cm2 at 4 h (p<0.001) and was inhibited completely (p<0.001) after 8 h by 100,000 MOI of bacteriophage BPECO19. At 10,000 and 1,000 MOI, initial contamination level of E. coli O157:H7 reduced to 3.10 Log CFU/cm2 (p<0.001) in 48 h and 4.56 Log CFU/cm2 (p<0.01) in 8 h, respectively (Fig. 3). After those time points, populations of E. coli O157:H7 slightly increased or maintained. However, it was clear that E. coli O157:H7 was significantly reduced by treatment of bacteriophage BPECO19 in beef.
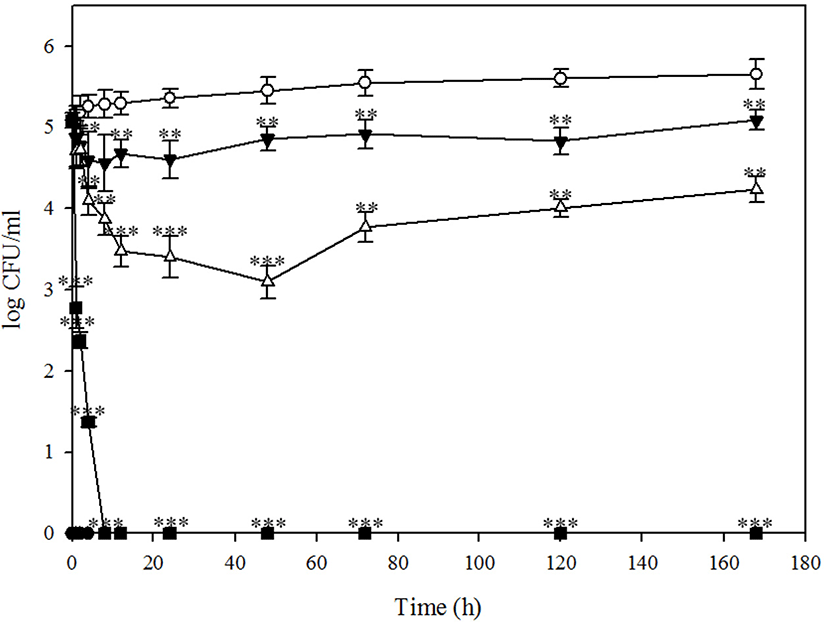
In comparison of storage temperature at 4℃ and 37℃, treatment of BPECO19 significantly reduced E. coli O157: H7 in pork at 4℃. Whereas 100,000 MOI of bacteriophage BPECO19 completely inhibited E. coli O157:H7 in beef in 8 h (p<0.001), E. coli O157:H7 spiked in pork was completely inhibited not only by 100,000 MOI (p<0.001) in 8 h but also by 10,000 MOI in 72 h (p<0.001) (Fig. 4). A significant 0.53 log (p<0.001) reduction of E. coli O157:H7 by 1,000 MOI of bacteriophage was observed after 8 h (Fig. 4). These findings demonstrate that inhibition of E. coli O157:H7 by bacteriophage BPECO 19 was more effective in pork than in beef (Fig. 4).
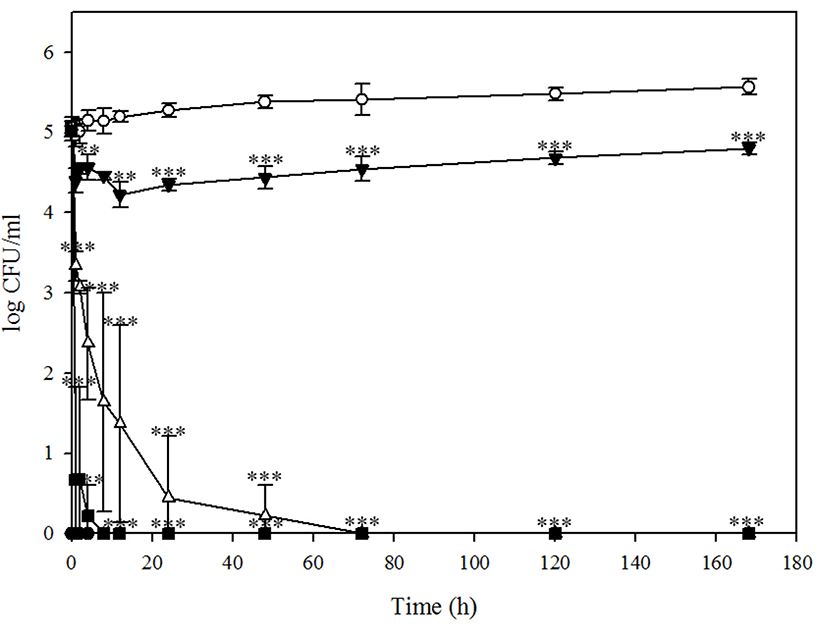
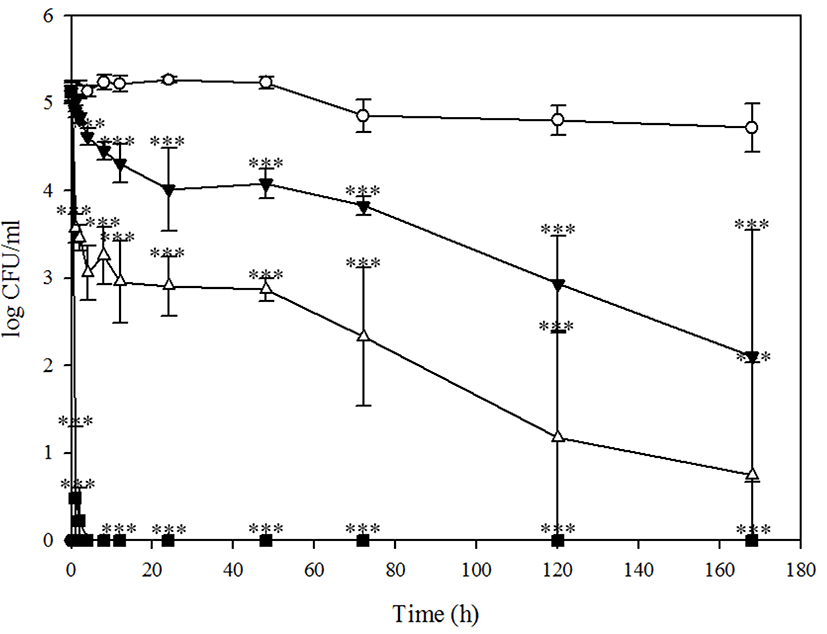
Although BPECO19 could not control E. coli O157:H7 in chicken stored at 37℃ (data not shown), it significantly reduced E. coli O157:H7 in chicken under the refrigeration condition. Whereas a transient increase in E. coli O 157:H7 bacterial load after bacteriophage BPECO19 treatment was observed in beef and pork, suppression of E. coli O157:H7 was constant in chicken meat (Fig. 5). A rapid 4.65 log reduction of E. coli O157:H7 by 100,000 MOI was reported after 1 h (p<0.001), a 4.90 log (p<0.001) reduction observed after 2 h and host bacteria was absent after 4 h (p<0.001) (Fig. 5). Whereas E. coli O157: H7 was not reduced completely by lower number of MOI, after 168 h, 1,000 and 10,000 MOI of bacteriophage BPECO19 significantly reduced E. coli O157:H7 by 3.02 log (p<0.001) and 4.39 log (p<0.001), demonstrating inhibition of E. coli O157:H7 in chicken meat.
Discussion
In this study, bacteriophage BPECO19 with prominent plague and lytic activity under in vitro conditions was the most promising strain against E. coli O157:H7 although all bacteriophage strains inhibited E. coli O157:H7 with various degrees. When bacteriophage BPECO19 was applied to control E. coli O157:H7 in artificially contaminated meat samples, it significantly reduced E. coli O157:H7 in beef, pork, and chicken stored at 4℃ which is used to keep fresh meats. Similarly, bacteriophage A511 and P100 significantly reduced L. monocytogenes in seafood, cheese, brine, chocolate milk, and vegetables under refrigerated conditions (Atterbury, 2009; Sabour and Griffiths, 2010). Because the growth of mesophilic bacteria is decreased or halt under refrigeration temperature (Cho et al., 2011), the resumption of E. coli O157:H7 did not exceed the lytic activity of bacteriophage BPECO19 in meat at 4℃. Combined with the lytic activity of bacteriophage BPECO19, cold temperature may contribute to effectively reduce E. coli O157:H7 in meat.
Compared with different temperatures, previous studies interestingly reduced pathogen at high temperature more than at low temperature or under refrigeration condition (Hudson et al., 2015; O'Flynn et al., 2004; Viazis et al., 2011a, 2011b). Their data was explained by the principle that diffusion or absorption rate of bacteriophage was proportionate to temperature. As bacteriophage adsorption was determined by second-order kinetics, −dp/dt=κPB (κ: phage absorption rate, P: phage concentration, B: bacterial concentration), high temperature seemed to increase the lytic activity of bacteriophage mixture BEC8 (Sabour and Griffiths, 2010; Viazis et al., 2011a, 2011b). On the contrary, the other study demonstrated that phage FAHEc1 was not effective to control E. coli O157:H7 in meat at 37℃ but in broth at 4℃ because the bacterial resumption at high temperature exceeded the lytic activity of bacteriophage (Hudson et al., 2013). Previous study was consistent with this finding that BPECO19 could not control E. coli O157:H7 in meat stored at 37℃ (Hudson et al., 2013). Therefore, storage at high temperature is not recommended to fresh meats because of the risk of outgrowth of spoilage bacteria or pathogens although some phages used in previous studies could be effective to control certain pathogens.
In accordance of food matrix or type of surface, MOI or concentration of bacteriophages varied in previous applications (Abuladze et al., 2008; Bigwood et al., 2008; Viazis et al., 2011a, 2011b; Worley-Morse et al., 2014). While 100 MOI of bacteriophage cocktail was effective to control pathogens on hard surfaces or leafy green vegetables, an MOI higher than 10,000 or 107 PFU/mL of bacteriophages was used in previous studies to control E. coli O157: H7 on cooked or raw beef (Hudson et al., 2013; Viazis et al., 2011a, 2011b). It was reported that 20,000 MOI of bacteriophage FAHEc1 decreased 1.4×104 CFU E. coli O157:H7 to undetectable level on beef in 1 h and lower MOI of FAHEc1 was not effective (Hudson et al., 2013). In other study to extend the shelf life of rib-eye steaks, 10,000 MOI of Pseudomonas bacteriophage could limit meat discoloration without sensory change, whereas low MOI could not (Greer, 2005). Like previous studies, lytic activity of bacteriophages BPECO19 significantly increased with MOI (Bigwood et al., 2008; Hudson et al., 2013; Hudson et al., 2015; Leverentz et al., 2003; O'Flynn et al., 2004; Sharma et al., 2009; Viazis et al., 2011a, 2011b; Worley-Morse et al., 2014). For the application of bacteriophage on meat, the higher than 10,000 MOI treatment of BPECO19 were needed to control E. coli O157:H7. As it is critical for food applications and medical treatments to determine the optimal MOI of bacteriophages, the preparation of high titer stocks in large volumes is required for bacteriophage applications in agriculture, animal husbandry, and food science (Hagens and Offerhaus, 2008; Worley-Morse et al., 2014). Since BPECO19 generally reached a titer of 1011 PFU/ml after 5 h of propagation at 37℃ (data not shown), this strain could be very useful for applications that require high titer bacteriophage.
Compared with meat samples, inhibition of E. coli O157: H7 by BPECO19 was prominent in beef and pork rather than chicken in this study. Several factors were suggested to inhibit the lytic activity of bacteriophage in meat and milk (O'Flynn et al., 2004; Sabour and Griffiths, 2010). Among them, the high moisture content of the meats reduced the lytic activity by reducing the diffusion of bacteriophages to the host bacteria (Sabour and Griffiths, 2010). As phage diffusion was determined by phage intrinsic properties and food matrix (Clokie and Kropinski, 2009), the moisture and properties of each meat samples might contribute to the difference in diffusion rate and lytic activity of bacteriophage. Similarly, the binding to heat-labile immunoglobulin proportionally reduced the lytic activity of phage K against Staphylococcus aureus (S. aureus). In addition, whey protein inhibited binding of phage K to S. aureus (Gill et al., 2006). In meat samples, abundance of immunoglobulins may affect the activity of bacteriophage BPECO19 and presence of additional inhibitory factors may explain the high titer required to inactivate host pathogen in meat samples.
The emergence of phage-resistant bacteria was a critical issue to be solved for the application of bacteriophages in food industry. Thus, a cocktail of broad-host-range phages was recommended in bacteriophage applications in order to prevent or reduce the chance of development of bacteriophage-resistant bacteria (Sabour and Griffiths, 2010). Three E. coli O157:H7 lytic bacteriophage cocktail ECP-100 is commercially called as EcoShield and was used to control E. coli O157:H7 in tomatoes, spinach, broccoli, lettuce, steak, and ground beef (Abuladze et al., 2008; Carter et al., 2012). The bacteriophage cocktail BEC8 also significantly reduced E. coli on steel and ceramic chips surfaces (Viazis et al., 2011a, 2011b).
However, several controversial studies reported that the use of bacteriophage cocktail could be associated with the emergence of bacteriophage-resistant bacteria (Abuladze et al., 2008; Bigwood et al., 2008; Worley-Morse et al., 2014). As the adsorption of bacteriophage to the cell wall of the host bacteria requires a specific receptor, competition for binding to the shared receptor by bacteriophages in cocktails could cause mutations of receptor against E. coli O157:H7 (Abuladze et al., 2008; O'Flaherty et al., 2005). Accordingly, the treatment with single bacteriophages such as Listeria monocytogenes or bacteriophage P- 100 was not associated with appearance of bacteriophage–resistant bacteria (Carlton et al., 2005; Viazis et al., 2011b; Worley-Morse et al., 2014). Therefore, it is of interest that bacteriophage BPECO19 alone successfully controlled E. coli O157:H7 in three types of meat samples. Considering that the emergence of bacteriophage-resistant or antibiotic-resistant bacteria might become a significant problem, the effect of single or cocktail treatment of bacteriophages on host mutation should be invested in further study.
To summarize, bacteriophage BPECO19 strain was selected for the biocontrol of E. coli O157:H7 under in vitro condition and on beef, pork, and chicken meat. In addition, bacteriophage BPECO19 alone reduced E. coli O157: H7 in a MOI-dependent manner under refrigerated condition. Although this study used the simple spreading method of bacteriophage at 100,000 MOI, the spraying method and combination with antimicrobials were attempted to enhance the biocontrol efficiency (Leverentz et al., 2003; Sharma et al., 2009). As an antimicrobial aerosol was successfully used to control foodborne pathogens in pork (Lee and Choi, 2012), aerosolization technique of bacteriophages could be applied to enhance meat safety in the further study.













