Introduction
Whey is the principal by-product of cheese manufacture. The huge quantities of whey produced by the dairy industry during cheese-making are difficult to dispose of, creating a serious environmental issue (Domingues et al., 1999; Marshall, 2004). However, whey contains a rich and heterogeneous mix of proteins, including β-lactoglobulin, α-lactalbumin, bovine serum albumin, glycomacropeptide, immunoglobulins, and lactoferrin (Walzem et al., 2002). Thus, this protein provides antimicrobial activity, immunomodulation, improved muscle strength, and lessens several risk factors for metabolic diseases (Bjørnshave and Hermansen, 2014; Marshall, 2004).
Bioconversion is known as a structural modification of chemical compounds (Asha and Vidyavathi, 2009). The recognition of bioconversion by microbes as an important tool has increased in the chemical, pharmaceutical, cosmetic, and food industries (Hegazy et al., 2015; Smitha et al., 2017). Lactic acid bacteria (LAB) have been traditionally used as the starter culture in fruits, vegetables, and dairy products, as well as probiotics (Sornplang and Piyadeatsoontorn, 2016). In addition, various enzymes, such as β-glucosidase, produced by LAB also utilize nutritional substances, providing benefits to the host (Lin et al., 2002). Furthermore, LAB associated with the bioconversion provide health-promoting attributes (Boguta et al., 2014; Lee and Paik, 2017).
Obesity is defined as the excessive accumulation of fat in the adipose tissue as a result of the imbalance between energy intake and expenditure (Spiegelman and Flier, 2001). The surplus energy derived from the metabolism of dietary carbohydrates, fats, and proteins leads to hypertrophic and hyperplasic adipose tissue expansion (Yang et al., 2019). The dramatic increasing incidence of metabolic diseases, including obesity, imposes a growing economic and medical burden on societies. Therefore, the biological components produced during whey bioconversion by LAB might be beneficial for metabolic diseases, such as obesity. In this study, we investigated whether bioconversion of whey using two LAB, Pediococcus pentosaceus and Lactobacillus sakei, reduces adipogenesis of preadipocytes.
Materials and Methods
Two LAB isolated from kimchi (P. pentosaceus KI 31 and L. sakei KI 36) were kindly provided by Dr. Sang-Dong Lim (Korea Food Research Institute, Wanju, Korea). Both LAB were cultured in de Man–Rogosa–Sharpe (MRS) medium (Neogen, Lansing, MI, USA) at 37°C. Whey bioconversion medium was prepared by adding 10% (w/v) whey powder (Samik Dairy & Food, Seoul, Korea) to distilled water, followed by pasteurization at 80°C for 1 min. The whey bioconversion medium was inoculated by either P. pentosaceus KI 31 or L. sakei KI 36 (1%) and incubated at 37°C for 24 h. After incubation, cell-free supernatants were collected by centrifugation and adjusted to pH 6.5 with 1N NaOH. The filtrate was obtained by passing the cell-free supernatants through a syringe filter. Protein concentrations of P. pentosaceus KI 31- and L. sakei KI 36-bioconverted whey (KI 31-W and KI 36-W, respectively) were measured using a bicinchoninic acid protein assay kit (Thermo Fisher Scientific, Rockford, IL, USA).
The mouse preadipocyte cell line, 3T3-L1, was purchased from the Korean Cell Line Bank (Seoul, Korea) and cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% bovine calf serum (BCS), 100 U/mL penicillin, and 100 μg/mL streptomycin (HyClone, Logan, UT, USA) at 37°C in humidified air containing 5% CO2. For adipocyte differentiation, 2-day post-confluent 3T3L-1 cells were stimulated with different concentrations of KI 31-W or KI 36-W in DMEM supplemented with 10% BCS, 0.5 mM 3-isobutyl-1-methylxanthine, 1 μM dexamethasone, and 5 μg/mL insulin (day 0). The cells were re-stimulated with KI 31-W or KI 36-W on day 3 and re-fed every 3 days with DMEM supplemented with 10% BCS and 5 μg/mL insulin. After incubation for 6 days, the medium was replaced with DMEM supplemented with 10% fetal bovine serum and antibiotics, and the cells were further incubated for 3 days.
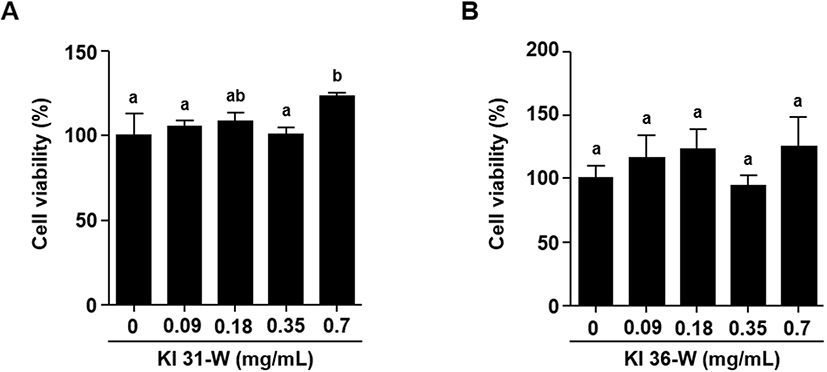
The 3T3L-1 cells were seeded on a 96-well culture plate and grown for 48 h. After the treatment with the different concentrations of KI 31-W or KI 36-W for 24 h, MTT (3-[4,5-dimethylthiazole-2-yl]-2,5-diphenyltetrazolium bromide) solution (0.5 mg/mL) was added to the medium and incubated at 37°C for 4 h. The resulting formazan dark purple crystal products were dissolved in 100 μL of dimethyl sulfoxide and the absorbance was measured at 570 nm using a microplate reader (Allsheng, Hangzhou, China).
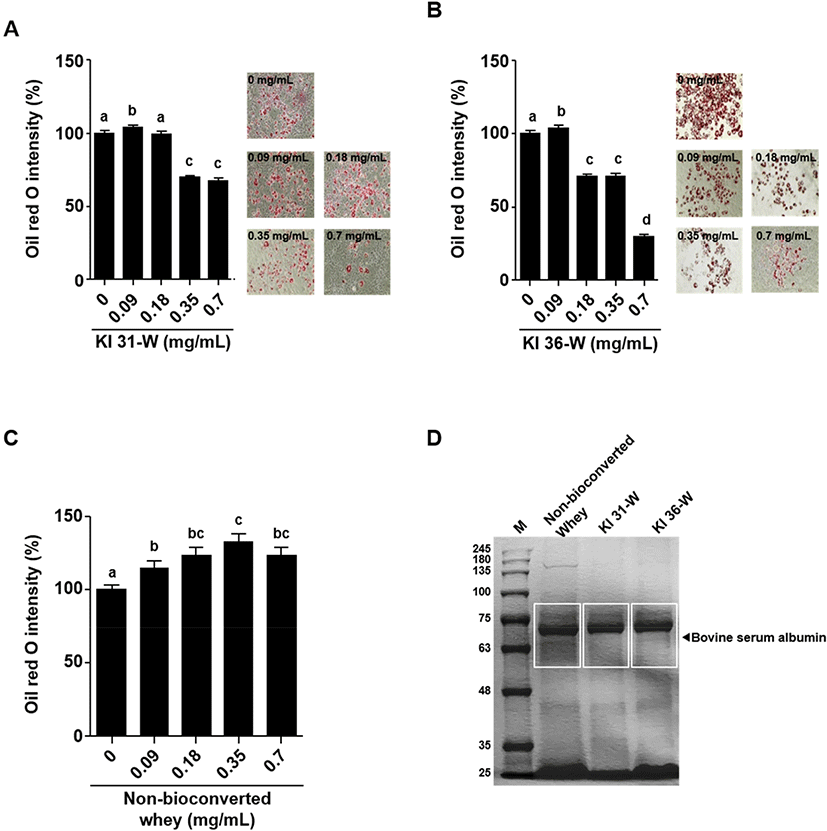
3T3-L1 cells treated with KI 31-W or KI 36-W were washed with phosphate-buffered saline and fixed with 10% formaldehyde for 1 h. After removing the formaldehyde, the cells were washed with 60% isopropanol and stained with Oil Red O solution (Sigma-Aldrich, St. Louis, MO, USA) for 30 min. Quantification of lipid accumulation was measured at 492 nm using a microplate reader (Allsheng). Microscopic images were also captured to visualize red oil droplet staining in differentiated 3T3-L1 cells treated with or without KI 31-W or KI 36-W. In addition, the lipid accumulation of 3T3-L1 cells treated by non-bioconverted whey was also determined by Oil Red O staining described above.
Non-bioconverted whey and bioconversion of whey by P. pentosaceus KI 31 or L. sakei KI 36 (15 μL each) were loaded on the gel and separated using 10% SDS-PAGE. The separated proteins were visualized by Coomassie blue staining.
After the differentiation of 3T3L-1 cells treated with the different concentrations of KI 31-W or KI 36-W, total RNA was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Then, complementary DNA was synthesized using total RNA (5 μg), random hexamers, and reverse transcriptase (Promega, Madison, WI, USA). Afterward, real-time PCR was performed with SYBR Green Real-Time PCR Master Mix (Toyobo, Osaka, Japan) using a Bio-Rad CFX thermal cycler (Bio-Rad Laboratories, Hercules, CA, USA). The following sequences of specific primers were used in this study. Peroxisome proliferator-activated receptor γ (PPARγ) (forward: 5’-AAGGGTGCCAGTTTCGATCC-3’, reverse: 5’-TCCTTGGCCCTCTG AGATGA-3’), adipocyte fatty acid-binding protein 2 (aP2) (forward: 5’- GCCCAACATGATCATCAGCG-3’, reverse: 5’-TGGTCGACTTTCCATCCCAC-3’), lipoprotein lipase (LPL) (forward: 5’-AACATTCCCTTCACCCTGCC-3’, reverse: 5’-GTCTCTCCGGCTTTCACTCG-3’), and β-actin (forward: 5’-TACAGCTTCACCACCACAGC-3’, reverse: 5’-GGAAAAGA GCCTCAGGGCAT-3’). After initial heating at 95°C for 10 s, 40 cycles of PCR (95°C for 5 s and 60°C for 31 s) were performed, followed by cooling at 4°C for 10 min. The relative expression level of genes was normalized to the β-actin level by using the 2−ΔΔCt method.
Results and Discussion
To examine the cell viability of 3T3-L1 cells in the presence of KI31-W or KI 36-W, the cells were treated with the indicated concentrations (0, 0.09, 0.18, 0.35, and 0.7 mg/mL) of KI 31-W or KI 36-W and the cell viability was determined using MTT assay. Fig. 1A and 1B indicated that bioconversion of whey by P. pentosaceus KI 31 or L. sakei KI 36 did not affect 3T3-L1 cell viability. Decreased lipid accumulation in mature 3T3-L1 cells in the presence of KI 31-W or KI 36-W was observed (Fig. 2A and 2B, respectively). A significantly reduced lipid accumulation was evident in 3T3-L1 cells treated with KI 31-W at 0.35 and 0.7 mg/mL (Fig. 2A). Likewise, lipid accumulation in 3T3-L1 cells was significantly inhibited by KI 36-W treatment at 0.18, 0.35, and 0.7 mg/mL. In particular, the lipid accumulation was reduced by over 50% following treatment with 0.7 mg/mL KI 36-W (Fig. 2B). However, non-bioconverted whey did not reduce lipid accumulation in 3T3-L1 cells (Fig. 2C). Although whey protein and its derivatives confer health benefits against several metabolic diseases (Patel, 2015), it has not been well established that whey protein prevents adipogenesis. Enriched whey protein fractions inhibit adipogenesis of 3T3-L1 (Rajic et al., 2010). More recently, D’Souza et al. (2020) demonstrated that whey protein hydrolyzed by pepsin and pancreatin enhances 3T3-L1 adipocyte differentiation and triacylglycerol accumulation. However, it has not yet been reported that bioconversion of whey exerts an anti-adipogenic effect. Our findings showed that bioconversion of whey by LAB effectively inhibits adipogenesis without decreasing the proliferation of 3T3-L1 cells. Further, our observation demonstrated that, among the proteins separated by SDS-PAGE, bovine serum albumin from bioconversion of whey by LAB was decreased compared to the protein of non-bioconverted whey (Fig. 2D). It has been shown that bovine serum, which contains albumin, facilitates the proliferation and differentiation of cells (Cho et al., 2018), assuming that the decreased bovine serum albumin by the bioconversion may be associated with the reduction of adipogenic differentiation.
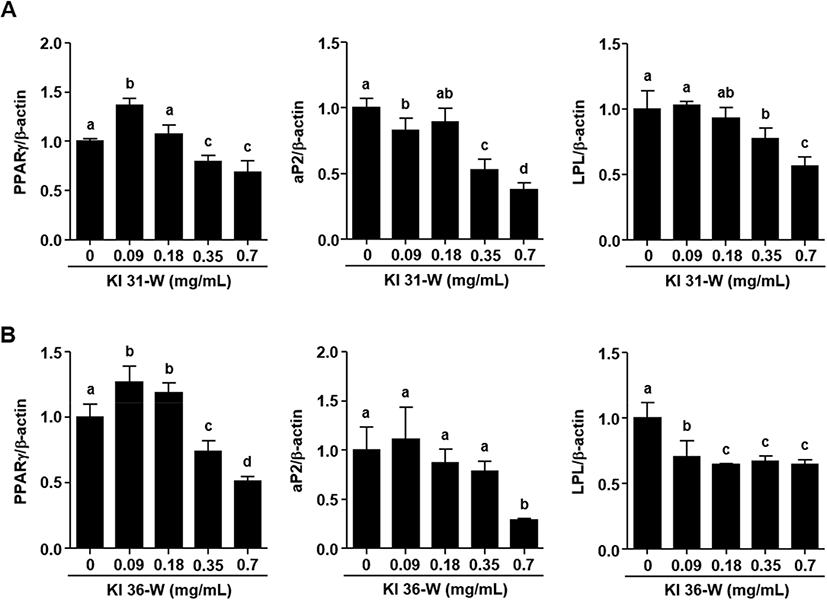
As shown in Fig 3A, KI 31-W remarkably decreased mRNA expression of PPARγ at doses of 0.35 and 0.7 mg/mL in 3T3-L1 cells. Similar effects on the down-regulation of aP2 and LPL were seen in the cells treated with 0.35 and 0.7 mg/mL. Fig. 3B demonstrated that KI 36-W also significantly down-regulated PPARγ at doses of 0.35 and 0.7 mg/mL in 3T3-L1 cells. However, aP2 mRNA expression was significantly decreased at 0.7 mg/mL KI36-W, but not at 0.35 mg/mL. In addition, LPL mRNA expression was strongly inhibited by KI 36-W at 0.09 to 0.7 mg/mL. Exposure of 3T3-L1 cells to dexamethasone, isobutylmethylxanthine, insulin, and fetal bovine serum, triggers a cascade of transcriptional activity, resulting in the rapid induction of regulators, such as PPARγ (Morrison and Farmer, 2000). Activation of PPARγ positively regulates downstream target genes, such as aP2 and LPL, associated with growth and differentiation (Farmer, 2005). Adipogenesis is strongly regulated by sequential activation of various transcription factors, such as PPARγ (Li et al., 2016). A dramatic increase in PPARγ gene expression was observed 5 days after the induction of 3T3-L1 cell differentiation (White and Stephens, 2010). The reduced expression of PPARγ was observed from an early stage (day 3) and final late-stage (day 8) during 3T3-L1 cell differentiation (Li et al., 2016). Our present study also showed that both KI 31-W and KI 36-W significantly suppressed PPARγ gene expression 9 days after the induction of differentiation. In addition, increased PPARγ expression sequentially up-regulates specific adipocyte genes, such as aP2 and LPL, to control fatty acid metabolism (Hassan et al., 2007). As a member of the fatty acid-binding proteins, aP2, which is highly expressed in mature adipocytes, plays an essential role in the intracellular fatty acid transport and metabolism critical for maintaining glucose and lipid homeostasis (Elmasri et al., 2009). LPL plays a pivotal role in the generation of fatty acids and their uptake by cells. Hence, the reduction of LPL expression during adipocyte differentiation may lead to decreased lipid storage in cells (Linehan et al., 2012).
In conclusion, we found that bioconversion of whey by LAB has the potential to suppress the adipogenic differentiation of 3T3-L1 cells, although it does not inhibit cell proliferation. Bioconversion of whey by LAB effectively and markedly reduces a key adipogenesis transcription factor, PPARγ, as well as specific lipogenesis genes, aP2 and LPL, consequently attenuating the adipogenic differentiation. To our best knowledge, this study is the first to demonstrate that bioconversion of whey has a beneficial effect.













