Introduction
Listeria monocytogenes is a gram-positive, facultative anaerobic bacterium that can proliferate at low temperatures (Walker et al., 1990) and survive in diverse environments, including NaCl concentrations up to 10% (McClure et al., 1989) and under acidic conditions (Cole et al., 1990). L. monocytogenes is a invasive bacterium, which is able to invade the human epithelial cells (Galdiero et al., 1997). In addition, L. monocytogenes is a pathogen that causes listeriosis, which is associated with septicemia, stillbirth, abortion, etc. (Gillespie et al., 2006). Listeriosis is usually linked to the consumption of raw milk, soft cheeses made from raw milk, smoked fish, and processed meat products (fermented sausages etc.), which are formulated with NaCl (Muhterem-Uyar et al., 2015; Samelis and Metaxopoulos, 1999).
NaCl is used to improve the flavor of processed products and to preserve food products by damaging the contaminating bacterial cells (Breslin and Beauchamp, 1997; Sofos, 1984). However, the NaCl concentrations used in foods may not be sufficient to inactivate pathogenic bacteria, and thus contributes to increased pathogenicity and resistance of bacteria to various stresses such as salt, acid, and heat (Bae et al., 2012; Garner et al., 2006; Jo et al., 2014). Phan-Thanh et al. (2000) found that L. monocytogenes adapted to an acidic environment (pH 5.2) for 2 h became resistant to heat and salt. In addition, NaCl-exposed E. coli O157:H7 NCCP11142 was heat resistant, and could survive at 50℃ (Lee et al., 2015). Also, Yoon et al. (2013) showed that heat resistance in Salmonella Typhimurium exposed to high NaCl concentration was increased.
Quantitative reverse trancription-PCR (qRT-PCR) was used to quantify the certain gene expression level (Livak and Schmittgen, 2001), and this method can be used to quantify gene expression levels by NaCl. For instance, Staphylococcus aureus upregulated the expression of genes related to biofilm formation when grown under high NaCl conditions (Rode et al., 2007).
To identify the invasive capability of L. monocytogenes, invasion assay using various human epithelial cell lines was usually performed, and the invasion efficiency was influenced by several stresses (Garner et al., 2006; Lee et al., 2013). Yoon et al. (2013) demonstrated that S. Typhimurium exposed to high NaCl concentration increased invasion efficiency into Caco-2 cells. In addition, Olesen et al. (2010) found that NaCl influences the invasiveness of L. monocytogenes.
Therefore, the objective of this study was to evaluate the effect of NaCl on the heat sensitivity of L. monocytogenes and to identify the genes expressed relatively in heatsensitized (HS) and non-sensitized (NS) strains to elucidate the correlation between NaCl and heat sensitivity in L. monocytogenes.
Materials and Methods
Nine L. monocytogenes strains (NCCP10805, NCCP 10806, NCCP10807, NCCP10808, NCCP10809, NCCP 10810, NCCP10811, NCCP10920, and NCCP10943), listed in Table 1 were individually cultured in 10 mL of tryptic soy broth containing 0.6% yeast extract (TSBYE; Becton, Dickinson, and Company, USA) at 30℃ for 24 h. Then, 0.1-mL aliquots of the cultures were transferred into 10 mL of fresh TSBYE and incubated at 30℃ for 24 h. The cultures were centrifuged (1,912 g, 15 min, 4℃), and the cells were washed twice with phosphate-buffered saline (PBS, pH 7.4; 0.2 g of KH2PO4, 1.5 g of Na2HPO4·7H2O, 8.0 g of NaCl, and 0.2 g of KCl in 1 L of distilled water) and then diluted with PBS to obtain 4 Log CFU/mL.
An aliquot (100 μL) of the inoculum was inoculated into 10 mL of TSBYE containing 0%, 1%, 2% and 4% NaCl, and incubated at 25℃ for 24-48 h. The cells were then plated on tryptic soy agar plus 0.6% yeast extract (TSAYE; Becton, Dickinson, and Company, USA) supplemented with 0%, 1%, 2%, and 4% NaCl and incubated at 25℃ for 48 h. After incubation, non-habituated L. monocytogenes (control) and NaCl-habituated L. monocytogenes cells (1-4%) growing on the plates were collected with a sterile bent glass rod, washed twice with PBS, and diluted with PBS to OD625=0.1. Then,1 mL aliquots of the L. monocytogenes strains were inoculated into 9 mL of TSBYE preheated to 60℃ in a water bath. To enumerate L. monocytogenes survival at 0, 20, 40 and 60 min, samples were removed at each time point, serially diluted with 0.1% buffered peptone water (BPW; Becton, Dickinson, and Company, USA), and plated on TSAYE. The plates were incubated at 30℃ for 48 h. L. monocytogenes survival was expressed as Log(Yt/Y0), where Yt is the cell count (Log CFU/mL) at time t and Y0 is the initial cell count (Log CFU/mL). Based on the heat challenge results, the nine tested L. monocytogenes strains were categorized as heat-sensitive (HS) or non-sensitive (NS).
To determine the relative expression levels of genes that were related to virulence, and osmotic and general stresses (inlA, inlB, opuC, betL, gbuB, osmC and ctc; Table 2) after exposure to NaCl, 0.4 mL of HS and NS L. monocytogenes inocula were inoculated into 40 mL of TSBYE and incubated at 25℃ to an OD625=0.6. After incubation, 9-mL aliquots of the cultures were exposed to TSBYE plus 0%, 1%, 2% and 4% NaCl for 20 min. Then, 1.5-mL aliquots of the cultures were placed in microtubes and centrifuged at 5,000 g for 5 min. Then, 0.1 mL of lysozyme (10 mg/mL; Wako Pure Chemical Industries, Ltd., Japan) was added to the cell pellets and mixed vigorously. The mixture was incubated at 37℃ for 15 min. After incubation, mRNA was extracted using the Qiagen RNeasy Mini Kit (Qiagen, Germany) and RNase-free DNase Set (Qiagen) according to the manufacturer’s instruction. Total mRNA was quantified by using an Epoch™ Microplate Spectrophotometer (BioTek Instruments, Inc., USA). The relative expression levels of virulence-, osmotic stress-, and general stress-related genes were measured by qRT-PCR. cDNA was synthesized from the extracted mRNA by using the QuantiTect Reverse Transcription Kit (Qiagen) according to the manufacturer’s instructions. The reaction mixture [24 μL; containing 12.5 μL of master mix, 6.5 μL of dH2O, and 2.5 μL of forward and reverse primers (10 pmol/μL)] was prepared by using the Rotor-Gene SYBR Green PCR Kit (Qiagen) according to the manufacturer’s protocol. Then, 1 μL of cDNA and 24 μL of the reaction mixture were added to a PCR strip. To determine the relative expression levels of the target genes, the data was analyzed using Rotor-Gene Q software (Qiagen). The mean threshold cycle (CT) values were used for the transcriptional analysis, and 16s rRNA was used as the reference gene to determine relative gene expression levels.
| Gene | Primer | Sequence (5′ → 3′) | Reference |
|---|---|---|---|
| 16s RNA | 16s RNA-F | CTA CGC ATT TCA CCG CTA CA | This study |
| 16s RNA-R | GAG GGT CAT TGG AAA CTG GA | This study | |
| inlA | inlA-F | GGT CTC ACA AAC AGA TCT AGA CCA AGT | Sue et al. (2004) |
| inlA-R | TCA AGT ATT CCA CTC CAT CGA TAG ATT | Sue et al. (2004) | |
| inlB | inlB-F | TGG GAG AGT AAC CCA ACC AC | This study |
| inlB-R | CGT CCC TGC CTC TAC TTT TG | This study | |
| opuC | opuC-F | CGG AAG ATC CCG TCA AAC TA | This study |
| opuC-R | CGT CAT ATG TGG CAT CAA GC | This study | |
| betL | betL-F | AAA CGA CAG GCG GAT CTT TA | This study |
| betL-R | CTT GCT ATC CCT GCT TGG AG | This study | |
| gbuB | gbuB-F | ATG ATG GCG GGT ATT AAC CA | This study |
| gbuB-R | CAT TGC ACC GAT CAT TGA AG | This study | |
| osmC | osmC-F | CTC CGT AAC CAG CAG CAA AT | This study |
| osmC-R | TCT CTG CAC CAA CAG AGC TT | This study | |
| ctc | ctc-F | CAG TTC GTG ACA ATG GTC GT | This study |
| ctc-R | CCT TTA ACG GGT CCA CTT GA | This study |
A Caco-2 cell invasion assay was performed to compare the invasion efficiency of the HS and NS L. monocytogenes strains according to the method by Lee et al. (2012).
Each experiment was tried twice with two samples per trial (n=4). The data for heat challenge and gene expression level were analyzed by the mixed model procedure and the general linear model procedure of SAS® (version 9.3; SAS Institute Inc., USA), respectively. A pairwise t-test at α=0.05 was used for all mean comparisons.
Results and Discussion
After heat challenge of the nine L. monocytogenes strains that were exposed to various NaCl concentrations, the strains were categorized in Table 3 as HS (NCCP10805, NCCP10806, NCCP10807, NCCP10810, NCCP10811, and NCCP10920) or NS (NCCP10808, NCCP10809, and NCCP10943). This result indicates that the cross-protective effect of NaCl on L. monocytogenes against heat is strain dependent. Therefore, it was necessary to find out what caused this strain variation. Palumbo et al. (1995) also showed that the survivability of L. monocytogenes in liquid egg yolk increased when 10% and 20% salt were added, because the D-value was higher as the temperature of the liquid egg yolk increased. In addition, the D-value of Salmonella spp. grown in liquid egg yolk containing 10% salt was higher than that in plain egg yolk (Palumbo et al., 1995). Lee et al. (2012) demonstrated that a mixture of 10 L. monocytogenes strains habituated by NaCl showed heat resistance, especially when they were exposed to sequentially higher NaCl concentrations (0%, 2%, 4%, and 6%). Kotrola and Conner (1997) showed that NaCl exposure increased the D-value of E. coli O157:H7 when grown at 52℃, 55℃, 57℃, and 60℃, indicating the increased survival of the bacterium. However, these studies did not identify the genes related to the cross-protection effect. Thus, we sought to analyze the gene expression levels of L. monocytogenes strains, exhibiting NaCl cross-protection to heat stress.
A-CDifferent letters in a same row mean significantly different at p<0.05.
In the HS strain NCCP10811, the relative expression levels of osmotic stress- and general stress-related genes (inlA, inlB, opuC and ctc) were not significantly increased by increasing NaCl concentrations (p>0.05) (Table 4; Fig. 1). Conversely, the relative expression levels of betL, gbuB and osmC, osmotic stress-related gene, increased as NaCl concentration increased (p<0.05) (Table 4; Fig. 1). However, in two of the NS strains (NCCP10808 and NCCP 10809), the relative expression levels of analyzed genes (inlA, inlB, opuC and ctc) were increased as the concentration of NaCl increased (p<0.05) (Table 4; Fig. 2). Osmotic- stress related genes are expressed as a response to osmotic stress conditions. In particular, inlA and inlB expression levels were much higher in the NS strains (p<0.05) than in the HS strain as the NaCl concentration increased. Therefore, the invasiveness of L. monocytogenes exposed to a high concentration of NaCl would likely increase. However, the invasion efficiency of the NS and HS L. monocytogenes strains in Caco-2 cells was not different (data not shown). Lee et al. (2012) also showed that exposure to NaCl did not affect human epithelial cell invasion of L. monocytogenes. These results indicate that there may be a threshold for inlA and inlB gene expression required for efficient L. monocytogenes invasion, and invasion efficiency may not be affected by inlA and inlB expression above the threshold. In other studies, the expression levels of betL, gbu and the opuC operon were increased as an adaptation to osmotic stress (Angelidis and Smith, 2003; Ko and Smith, 1999). Bae et al. (2012) showed that several transporters associated with the uptake of glycine and betaine were upregulated at 1.2% NaCl, which is a salt concentration commonly used in many RTE foods. In addition, the accumulation of inlA, opuC and opuA increased within 5 min when L. monocytogenes was exposed to osmotic stress when compared to the levels in the control, which was not exposed to osmotic stress (Sue et al., 2004). A study by Gardan et al. (2003) showed that ctc, a L. monocytogenes gene related to general stress, was expressed at higher levels under high osmolarity conditions when there were no osmoprotectants, including glycine and betaine. Duche et al. (2002) showed that salt shock proteins (Ssp) in L. monocytogenes rapidly increased after exposure to salt stress, and Ssp overexpression was retained several hours after shifting back to normal conditions.
A-CDifferent letters in a same column mean significantly different at p<0.05.
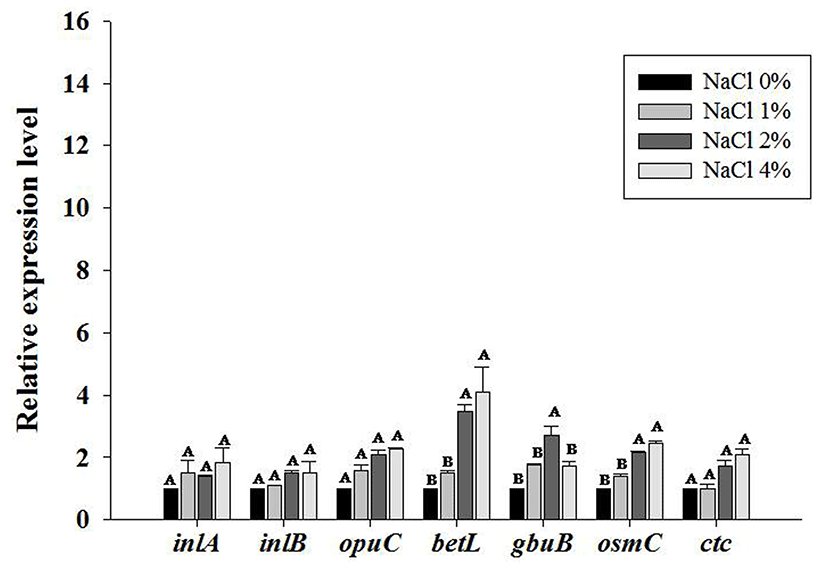
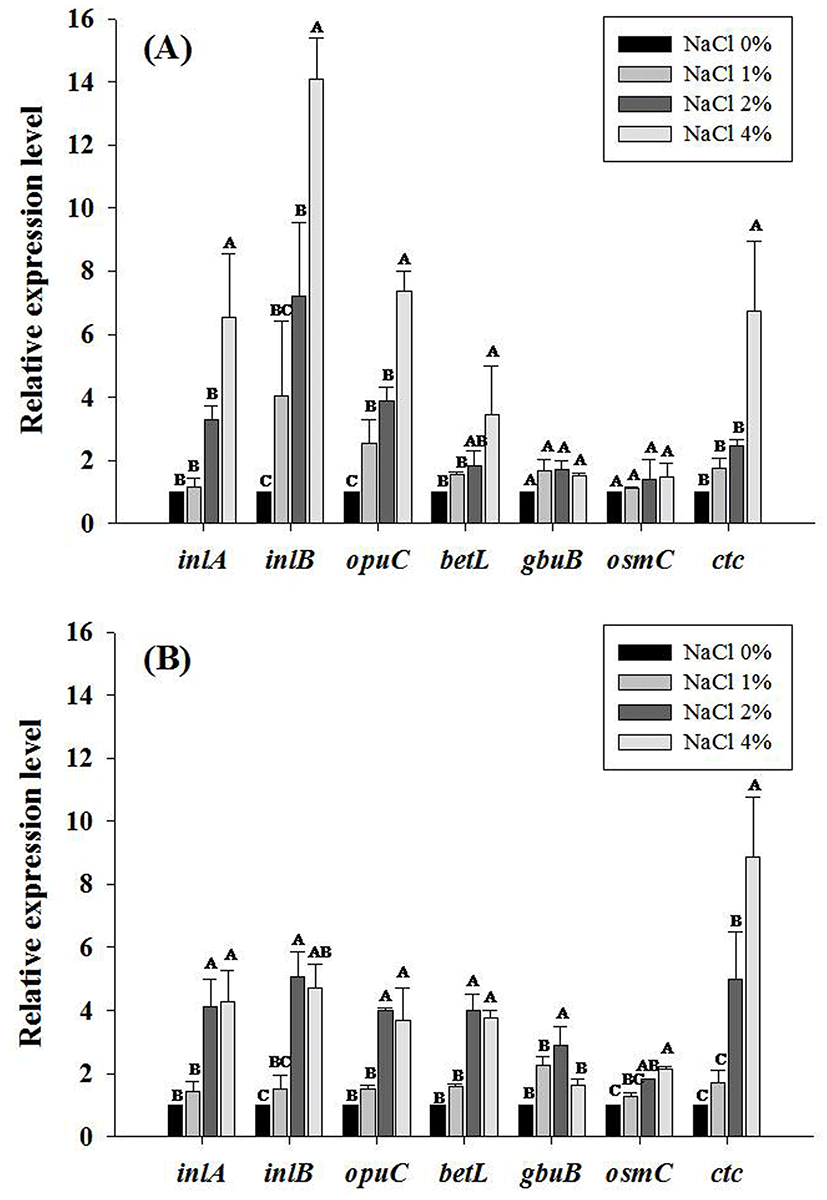
In conclusion, the effect of NaCl on heat-sensitization of L. monocytogenes is strain-dependent, and opuC and ctc may play a role in preventing heat-sensitization by NaCl in NS L. monocytogenes strains. In addition, NaCl exposure also increased the expression of invasion-related genes (inlA and inlB) in NS L. monocytogenes.













