Introduction
Probiotics, which mostly consist of lactic acid bacteria, are well known to confer beneficial effects (Chugh and Kamal-Eldin, 2020). Their physiological functions are beneficial to human health by inhibiting pathogen adhesion, enhancing the functions of the intestinal epithelial barrier, and modulating host immune responses (Lebeer et al., 2010). Moreover, the use of probiotics is a promising therapeutic approach against many inflammatory diseases (Maekawa and Hajishengallis, 2014). These effects of probiotics can be attributed to the production of bioactive molecules, such as cell wall components, bacteriocins, short-chain fatty acids, exopolysaccharides, enzymes, and amino acids (Chugh and Kamal-Eldin, 2020; Lebeer et al., 2010).
In addition to the bioactive molecules of probiotics, their genomic DNA (gDNA) has been proven to exert probiotic effects. For instance, gDNA extracted from Lactobacillus plantarum inhibited lipopolysaccharide (LPS)-induced tumor necrosis factor (TNF)-α secretion by suppressing the activation of mitogen-activated protein kinases (MAPKs) and nuclear factor-κB (NF-κB) in THP-1 cells (Kim et al., 2012). Moreover, oligodeoxynucleotides from L. casei gDNA inhibited inflammatory responses in human intestinal epithelial cells in vitro as well as in a dextran sulfate sodium-induced murine colitis model (Hiramatsu et al., 2014). More recently, gDNA extracted from L. rhamnosus GG was found to inhibit LPS-induced interleukin (IL)-6 production in RAW 264.7 cells (Qi et al., 2020).
Periodontitis is defined as a chronic inflammatory oral disease caused by multiple microbial infections, resulting in the gradual destruction of tissues supporting teeth and the alveolar bone (Saini et al., 2009). Moreover, periodontitis is known to be associated with systemic diseases, such as cardiovascular and respiratory diseases, rheumatoid arthritis, and metabolic syndrome (Winning and Linden, 2015). Porphyromonas gingivalis is a Gram-negative anaerobic bacterium that is considered to be the main pathogen responsible for periodontitis by producing several virulence factors, such as extracellular proteases and LPS (Bostanci and Belibasakis, 2012). Among the bacteria associated with the pathogenesis of periodontitis, P. gingivalis has been extensively studied because of its unique ability to penetrate the gingivae and cause tissue destruction by inducing inflammation (How et al., 2016).
The findings of in vitro and in vivo analyses have confirmed that probiotics can alleviate periodontal diseases (Nguyen et al., 2021). However, probiotics need to be viable to perform their biological functions. Moreover, the use of probiotics may cause some adverse effects, such as antibiotic resistance and systemic infections in immunocompromised patients (Doron and Snydman, 2015). Therefore, molecules with functional bioactivities, also known as postbiotics, are considered promising alternatives to probiotics. Although the health benefits of probiotics, particularly lactobacilli, are well known, pediococci are also exhibit certain probiotic effects. Bioactive molecules, such as exopolysaccharides and bacteriocins, of Pediococcus acidilactici exhibit probiotic effects, including antioxidant, lipid-lowering, antimicrobial, and anti-inflammatory effects (Moon et al., 2014; Seo and Kang, 2020; Song et al., 2021). In addition, our studies have shown the beneficial effects of postbiotics derived from P. acidilactici K10 and P. acidilactici HW01, such as bacteriocin and bacterial lysates. Bacteriocins of P. acidilactici K10 and P. acidilactici HW01 inhibit biofilm formation of foodborne pathogenic bacteria, including Enterococcus faecalis and Pseudomonas aeruginosa (Lee et al., 2020; Yoon and Kang, 2020). More recently, bacterial lysates of both P. acidilactici K10 and P. acidilactici HW01 exert antibiofim, anti-adhesive and anti-invasive activities against Listeria monocytogenes (Lee et al., 2022). Therefore, this study aimed to assess the anti-inflammatory effect of gDNA extracted from P. acidilactici K10 and P. acidilactici HW01 against P. gingivalis LPS-induced inflammatory responses.
Materials and Methods
P. acidilactici K10 and P. acidilactici HW01, which were isolated from kimchi and malt (Ahn et al., 2017; Kwon et al., 2002), were maintained in Man Rogosa Sharpe medium (Neogen, Lansing, MI, USA) at 37°C. The gDNAs of P. acidilactici K10 and P. acidilactici HW01 (K10 gDNA and HW01 gDNA, respectively) were extracted using the Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA) according to the manufacturer’s instructions, with some modifications. In brief, bacterial pellets were harvested by centrifugation at 13,000×g for 10 min and extensively washed with phosphate-buffered saline (PBS). The bacterial pellets were resuspended in 480 μL of 50 mM EDTA and 120 μL of lysozyme (10 mg/mL; Sigma-Aldrich, St. Louis, MO, USA) and incubated at 37°C for 30 min. After centrifuging at 13,000×g for 10 min, nuclei lysis solution (600 μL) was added to the pellets and incubated at 80°C for 5 min to lyse the cell membrane. RNase solution (3 μL) was then added to the lysates, which were mixed by inverting the tubes five times and then incubated at 37°C for 40 min. Protein precipitation solution (200 μL) was added to the mixture and incubated on ice for 5 min to purify the gDNA. After centrifuging at 13,000×g for 3 min, the supernatants were transferred to a clean microtube and gently mixed with 600 μL of isopropanol. After centrifuging at 13,000×g for 3 min, the supernatants were removed and the remaining pellets were washed with 70% ethanol. Finally, the ethanol was discarded and the pellets containing gDNA were rehydrated by adding DNA rehydration solution. The concentration and purity of gDNA were determined using a NanoDrop spectrophotometer (ND-1000, NanoDrop Technologies, Hampton, NH, USA).
RAW 264.7 cells were obtained from the American Type Culture Collection (ATCC; Manassas, VA, USA) and maintained in Dulbecco’s modified Eagle’s medium (DMEM; Welgene, Gyeongsan, Korea) containing 10% heat-inactivated fetal bovine serum (Gibco, Burlington, ON, Canada), 100 U/mL penicillin, and 100 μg/mL streptomycin (HyClone, Logan, UT, USA) at 37°C in a 5% CO2-humidified incubator.
RAW 264.7 cells (1×105 cells/mL) were plated on a 12-well culture plate and incubated at 37°C for 24 h. The cells were pretreated with K10 gDNA or HW01 gDNA (1 μg/mL) for 3, 9, or 15 h and stimulated with 1 μg/mL of P. gingivalis LPS for 3 h. In a separate experiment, RAW 264.7 cells (1×105 cells/mL) were plated on a 12-well culture plate and incubated at 37°C for 24 h. The cells were pretreated with various concentrations of K10 gDNA or HW01 gDNA (0.1, 1, and 10 μg/mL) for 15 h and stimulated with 1 μg/mL of P. gingivalis LPS for 3 h. Subsequently, spent culture supernatants were discarded and total RNA was extracted from the cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA), according to the manufacturer’s instructions. Total RNA was reversed transcribed to complementary DNA (cDNA) using random hexamers and reverse transcriptase (Promega). cDNA was amplified by qPCR with SYBR Green Real-Time PCR Master Mix (Toyobo, Osaka, Japan) using the StepOnePlusTM Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). Specific genes responsible for inflammatory responses, namely those encoding IL-1β, IL-6, and monocyte chemoattractant protein (MCP)-1, were amplified under the following conditions: Denaturation at 95°C for 10 s, amplification for 40 cycles at 95°C for 5 s and 60°C for 31 s. The sequences of specific primers were as follows: IL-1β, forward, 5′-CTCACAAGCAGAGCACAAGC-3′ and reverse, 5′-TCTTGGCCGAGGACTAAGGA-3′; IL-6, forward, 5′-TCCTACCCCAATTTCCAATGCT-3′ and reverse, 5′-TCTGACCACAGTGAGGAATGTC-3′; MCP-1, forward, 5′-AGCCAACTCTCACTGAAGCC-3′ and reverse, 5′-TCTCCAG CCTACTCATTGGGA-3′; and β-actin, forward, 5′-TACAGCTTCACCACCACAGC-3′ and reverse, 5′-GGAAAAGAGCCT CAGGGCAT-3′. The relative mRNA expression levels of IL-1β, IL-6, and MCP-1 were normalized to that of β-actin using the 2−ΔΔCt method.
RAW 264.7 cells (1×105 cells/mL) were plated on a 96-well culture plate and incubated at 37°C for 24 h. The cells were pretreated with 1 μg/mL of K10 gDNA or HW01 gDNA at 37°C for 15 h and stimulated with P. gingivalis LPS (1 μg/mL) at 37°C for further 24 h. Subsequently, spent culture supernatants were collected and the secretion levels of IL-1β, IL-6, and MCP-1 were measured using commercial ELISA kits (R&D Systems, Minneapolis, MN, USA), according to the manufacturer’s instructions.
RAW 264.7 cells (1×105 cells/mL) were plated on a 6-well culture plate and incubated at 37°C for 24 h. The cells were pretreated with 1 μg/mL of K10 gDNA or HW01 gDNA at 37°C for 15 h and stimulated with P. gingivalis LPS (1 μg/mL) at 37°C for 30 min. The cells were then washed with PBS and lysed with a lysis buffer (1 M HEPES, 1 M NaCl, 1% IGEPAL®-CA 630, 0.75% sodium deoxycholate, and 10% glycerol) containing protease and phosphatase inhibitors. Equal amounts of proteins were separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and electrotransferred to polyvinylidene difluoride membranes (Millipore, Bedford, MA, USA). The membranes were then blocked with 5% skimmed milk in Tris-buffered saline containing 0.1% Tween 20 (TBST) and incubated with primary antibodies specific to extracellular signal-regulated kinase (ERK), phosphorylated ERK, p38 kinase, phosphorylated p38 kinase, c-Jun-N-terminal kinase (JNK), phosphorylated JNK, IκBα (Cell Signaling Technology, Danvers, MA, USA), or β-actin (Santa Cruz Biotechnology, Santa Cruz, CA, USA) at 4°C for 15 h. The membranes were washed with TBST and then incubated with horseradish peroxidase-conjugated anti-rabbit IgG (Cell Signaling Technology) at room temperature for 1 h. Immunoreactive proteins were detected using an enhanced chemiluminescence reagent (Dyne Bio, Seongnam, Korea) and visualized using a C-DiGit blot scanner (Li-Cor Bioscience, Lincoln, NE, USA).
All experiments were performed thrice. The results are expressed as the means±SDs. Statistical differences between the appropriate control and gDNA-treated groups were assessed by performing unpaired two-tailed t-tests using GraphPad Prism 5 software (GraphPad Software, La Jolla, CA, USA).
Results
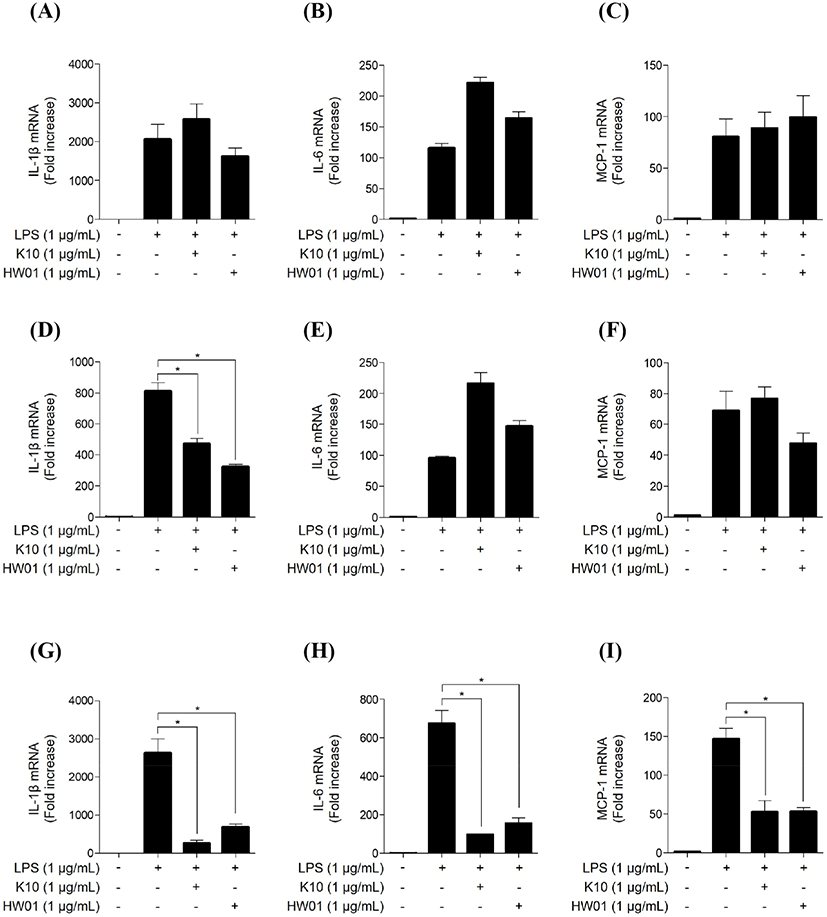
When the cells were pretreated with K10 gDNA and HW01 gDNA for 3 h, the gDNAs did not inhibit P. gingivalis-induced mRNA expression of IL-1β (Fig. 1A), IL-6 (Fig. 1B), and MCP-1 (Fig. 1C). However, when the cells were pretreated with K10 gDNA and HW01 gDNA for 9 h, the gDNAs partially suppressed inflammatory responses. As shown in Fig. 1D, P. gingivalis LPS-induced IL-1β mRNA expression was significantly downregulated in the presence of K10 gDNA and HW01 gDNA (p<0.05), whereas IL-6 and MCP-1 mRNA expression was not inhibited by these gDNAs (Figs. 1E and 1F, respectively). When RAW 264.7 cells were pretreated with gDNAs for 15 h, the gDNAs significantly inhibited the mRNA expression of IL-1β (Fig. 1G), IL-6 (Fig. 1H), and MCP-1 (Fig. 1I; p<0.05). As 15 h-pretreatment with gDNAs effectively inhibited P. gingivalis LPS-induced inflammatory responses, all subsequent experiments were performed using cells pretreated with gDNAs for 15 h.
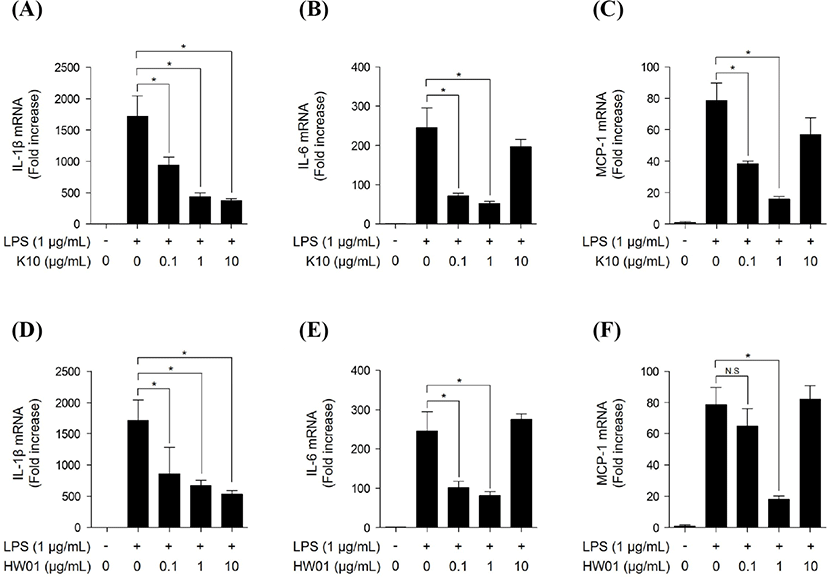
To assess the inhibitory effect of different concentrations of gDNAs on P. gingivalis LPS-induced inflammatory responses in RAW 264.7 cells, the cells were pretreated with gDNAs (0.1, 1, and 10 μg/mL) for 15 h and then stimulated with P. gingivalis LPS (1 μg/mL) for 3 h. As expected, P. gingivalis LPS alone induced IL-1β mRNA expression. However, both K10 gDNA and HW01 gDNA dose-dependently inhibited P. gingivalis LPS-induced IL-1β mRNA expression (Figs. 2A and 2D, respectively). Pretreatment with 0.1 and 1 μg/mL of K10 gDNA and HW01 gDNA significantly inhibited P. gingivalis LPS-induced IL-6 mRNA expression (p<0.05; Figs. 2B and 2E, respectively). However, 10 μg/mL of both gDNAs failed to inhibit P. gingivalis LPS-induced IL-6 mRNA expression (Figs. 2B and 2E). Similarly, 0.1 and 1 μg/mL of K10 gDNA significantly inhibited P. gingivalis LPS-induced MCP-1 mRNA expression, whereas 10 μg/mL did not (Fig. 2C). Moreover, 1 μg/mL of HW01 gDNA inhibited P. gingivalis LPS-induced MCP-1 expression, whereas 0.1 and 10 μg/mL did not (Fig. 2F). Therefore, we decided that the optimal concentration of gDNAs to inhibit P. gingivalis LPS-induced inflammatory responses was 1 μg/mL and used this concentration for further analysis.
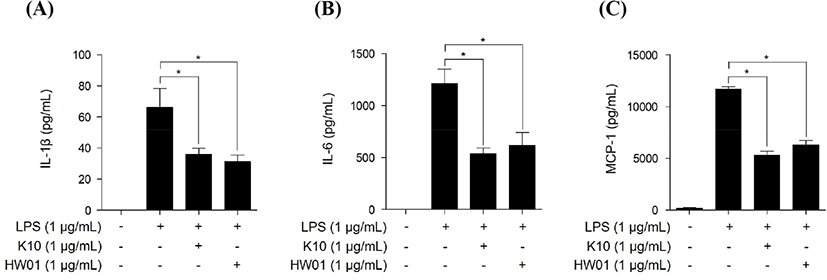
To further confirm the inhibition of inflammatory responses at the protein level, RAW 264.7 cells were pretreated with gDNAs (1 μg/mL) for 15 h and then stimulated with P. gingivalis LPS (1 μg/mL) for 24 h. Consistent with the results of mRNA expression, both gDNAs significantly inhibited IL-1β (Fig. 3A), IL-6 (Fig. 3B), and MCP-1 (Fig. 3C) secretion induced by P. gingivalis LPS (p<0.05). These results indicate that K10 gDNA and HW01 gDNA effectively inhibit P. gingivalis LPS-induced inflammatory responses.
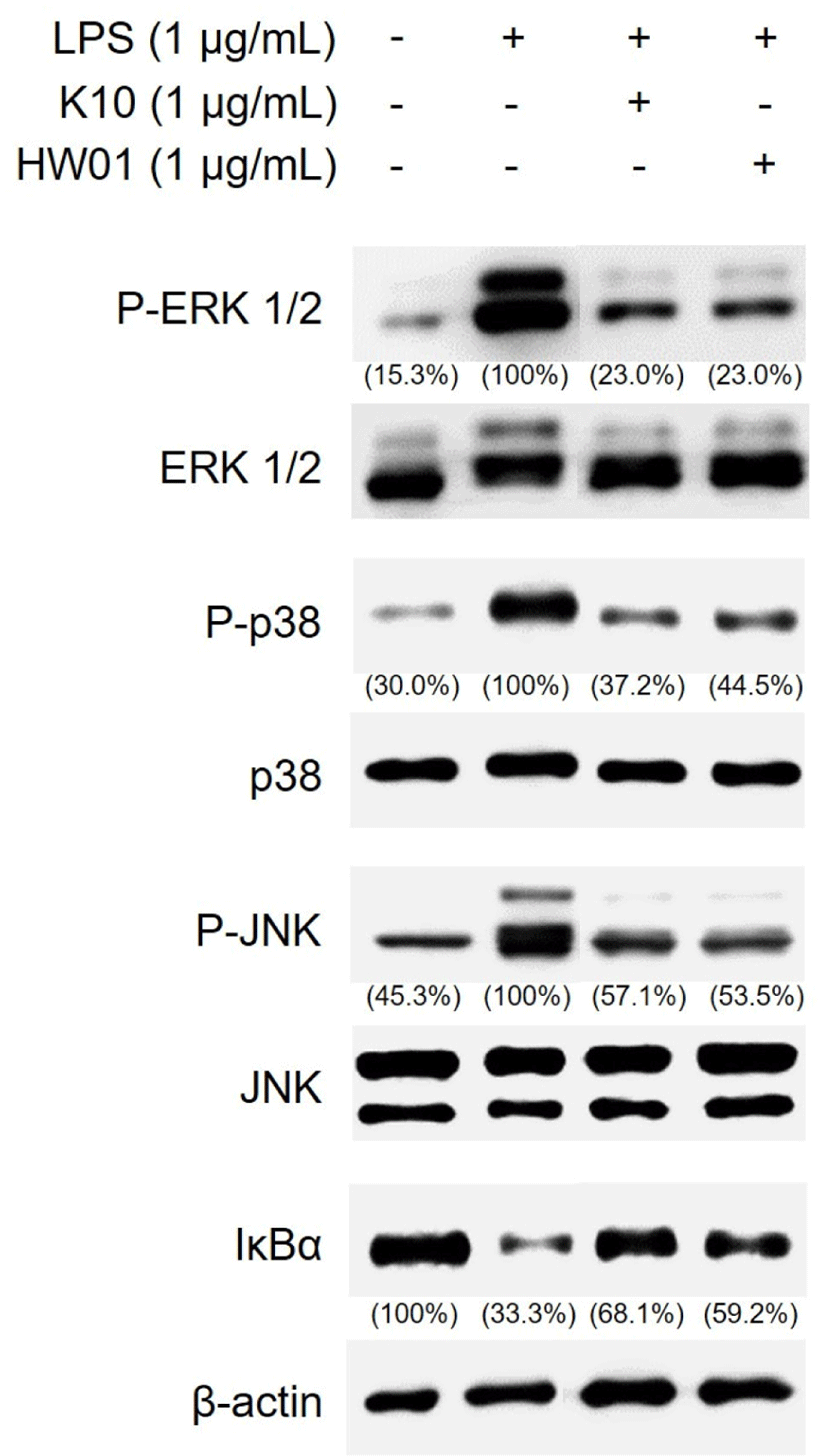
MAPK pathways, such as ERK 1/2, p38 kinase, JNK, and NF-κB pathways, are known to be common inflammatory signaling pathways that cause the release of proinflammatory cytokines, such as IL-1β, and consequently lead to inflammation (Jayakumar et al., 2021). Therefore, the activation of MAPK and NF-κB was assessed in RAW 264.7 cells pretreated with or without gDNAs. As shown in Fig. 4, the phosphorylation of ERK 1/2, p38 kinase, and JNK was higher in RAW 264.7 cells stimulated with P. gingivalis LPS than in those not treated with P. gingivalis LPS. However, pretreatment with K10 gDNA and HW01 gDNA abrogated the phosphorylation of ERK 1/2, p38 kinase, and JNK, indicating that ERK 1/2, p38 kinase, and JNK signaling pathways were blocked on pretreatment with both these gDNAs and consequently inhibited P. gingivalis LPS-induced inflammatory responses. Moreover, inhibitors of NF-κB, such as IκBα, are associated with the downregulation of NF-κB (Wang et al., 2020). P. gingivalis LPS alone inhibited IκBα degradation, indicating that the inflammatory responses were augmented by NF-κB activation. However, pretreatment of RAW 264.7 cells with K10 gDNA or HW01 gDNA (Fig. 4) restored IκBα degradation, suggesting that both these gDNAs attenuated NF-κB activation, which was associated with P. gingivalis LPS-induced inflammatory responses.
Discussion
Periodontitis is the most prevalent oral disease that is characterized by gingival inflammation and the loss of connective tissue attachment, resulting in alveolar bone resorption and tooth loss (Xu et al., 2020). Although the role of probiotics in modulating periodontal diseases is not fully understood, several studies have reported the protective effects of probiotics, particularly Lactobacillus spp., on periodontal diseases based on in vitro and in vivo analyses (Gatej et al., 2018; Nguyen et al., 2021). P. acidilactici is frequently found in fermented vegetables and meat-based products and is considered a potential probiotic with antimicrobial, antioxidant, and immunomodulatory effects (Feng et al., 2016; Papagianni and Anastasiadou, 2009; Song et al., 2021). Furthermore, the present study demonstrated that K10 gDNA and HW01 gDNA markedly inhibited P. gingivalis LPS-induced inflammatory responses by suppressing MAPK and NF-κB signaling pathways in RAW 264.7 cells, suggesting that P. acidilactici could also be a potential probiotic that could be beneficial to oral health.
Several lactobacilli have been reported to inhibit inflammatory responses induced by periodontal pathogenic bacteria, such as P. gingivalis. Co-culture with P. gingivalis and L. acidophilus significantly decreased the mRNA and protein expression levels of IL-1β, IL-6, and IL-8 in gingival epithelial cells in a dose-dependent manner (Zhao et al., 2012). More recently, mono-infection of gingival epithelial cells with P. gingivalis was found to increase the expression of IL-1β and TNF-α. However, treatment with L. rhamnosus Lr-32 or L. acidophilus LA-5 markedly decreased the expression of IL-1β and TNF-α (Albuquerque‐Souza et al., 2019). In fact, P. acidilactici has displayed anti-inflammatory activities against various pathogenic bacteria. We previously found that bacteriocins isolated from P. acidilactici K10 and HW01 suppressed IL-8 production in the human intestinal epithelial cell line Caco-2 in response to Enterococcus faecalis infection (Yoon and Kang, 2020). Cell-free supernatants of P. acidilactici TMAB26 effectively inhibited Klebsiella pneumoniae LPS-induced IL-6 and TNF-α mRNA expression in HT-29 cells and peripheral blood mononuclear cells (Barigela and Bhukya, 2021). However, the antagonistic effect of P. acidilactici or its component(s) on P. gingivalis-induced inflammatory responses has not been elucidated. In addition, gDNA derived from Lactobacillus spp. has been found to have anti-inflammatory properties. For example, high-frequency oligodeoxynucleotides of L. casei gDNA inhibited IL-8 production in Caco-2 cells in vitro and alleviated dextran sulfate sodium-induced murine colitis (Hiramatsu et al., 2014). Consistent with the present findings, pretreatment with L. rhamnosus GG gDNA was found to alleviate mRNA expression levels of inflammatory cytokines in RAW 264.7 cells upon LPS challenge (Qi et al., 2020). However, whether gDNAs of probiotic strains exhibit anti-inflammatory effects against oral pathogenic bacteria remains unclear. We found that P. acidilactici gDNA can inhibit P. gingivalis LPS-induced inflammatory responses.
During the development of periodontitis, periodontal pathogens trigger innate immune responses by releasing various cytokines and chemokines, which play a pivotal role in the progression of inflammation (Ramadan et al., 2020). Excessive IL-6 production may be associated with periodontal tissue destruction by inducing matrix metalloproteinase (MMP)-1 secretion during the progression of periodontitis (Sundararaj et al., 2009). In addition, IL-1β is involved in the production of MMP-1, leading to tissue damage (Cheng et al., 2020). MCP-1 is produced by macrophages and fibroblasts and acts as a chemoattractant for monocytes and neutrophils (Strecker et al., 2011). Excessive neutrophil activation can result in the production of toxic compounds and reactive oxygen species that can also affect aggressive periodontitis (Ramadan et al., 2020). We found that both K10 gDNA and HW01 gDNA significantly suppressed these cytokines/chemokines by regulating MAPK and NF-κB signaling pathways, suggesting that they could relieve inflammation and consequently prevent periodontitis. Although the precise mechanisms by which P. acidilactici gDNAs inhibit the production of cytokines/ chemokines responsible for periodontitis are not fully understood, it can be speculated that cell signaling mediated by toll-like receptor 9 (TLR9) negatively regulates inflammatory responses. TLR9 is a critical receptor for recognizing bacterial DNA (Chockalingam et al., 2012). Li et al. (2017) reported that TLR9 deficiency enhanced the activation of NF-κB, resulting in increased IL-1β expression in intestinal epithelial cells. Moreover, the use of CpG oligodeoxynucleotides as a TLR9 ligand inhibited MCP-1 expression in adipocytes (Thomalla et al., 2019). Although TLR signaling pathways mainly induce inflammatory responses, TLR9 signaling promotes the accumulation of IκBα in the cytoplasm, thereby suppressing NF-κB activation (Li et al., 2017). Therefore, TLR9 activation by P. acidilactici gDNAs may negatively regulate intracellular signaling pathways, such as MAPK and NF-κB pathways, thereby attenuating inflammatory responses.
Conclusion
In conclusion, the present study demonstrated the inhibitory effect of P. acidilactici gDNA on P. gingivalis LPS-induced inflammatory responses. Several probiotic lactobacilli have shown anti-inflammatory activities against P. gingivalis. Our results provide important insights into the inhibition of periodontal inflammation using P. acidilactici gDNA. Thus, P. acidilactici gDNA could be a potential therapeutic option against periodontitis.













