Introduction
Traditional fermented food products are prepared with biotechnological methods using indigenous microorganisms present in food (Utama et al., 2019). These methods are practical and economical to preserve food (Eze et al., 2014). Indonesia is home to traditional fermented food products such as bekasam, the naturally fermented fish from Central Java, South Sumatra, and Central Kalimantan. Today, bekasam is also made of rabbit meat.
Indonesians people generally avoid consuming rabbit meat because they consider rabbit as a pet. Therefore, rabbit meat is not widely distributed in Indonesian markets (Priyanti and Raharjo, 2012). Interestingly, the nutritional quality of rabbit meat is superior to other species due to high proteins, high biological value, minerals, and vitamins despite the low saturated fatty acids, cholesterol, and sodium (El-Medany and El-Reffaei, 2015; Nistor et al., 2013). Accordingly, rabbit meat-based products are recently developed to implement food diversification and to increase the consumption of rabbit meat in Indonesia.
Three key steps in preparing rabbit meat bekasam include salting process, carbohydrate addition (rice), and fermentation process. The salting process aims to select microbes and prevent the growth of pathogen bacteria as spoilage microorganisms (Doyle et al., 2001). Then, rice as the source of carbohydrate is added to stimulate the growth of lactic acid bacteria (LAB) (Putri et al., 2015). On fermentation, LAB breaks down carbohydrates into lactic acid, propionic acid, acetic acid, and ethyl alcohol (Ahmed et al., 2013). These compounds are useful preservatives and impart sour taste to bekasam (Anihouvi et al., 2012).
LAB is the major component in bekasam fermentation. Previous study on eight types of fish bekasam from eight areas in Indonesia reported 62 LAB isolates including 19 which exhibited antimicrobial activity against Escherichia coli, Salmonella typhimurium ATCC 14028, Bacillus cereus, Staphylococcus aureus, and Listeria monocytogenes (Desniar et al., 2013). Therefore, LAB is the potential inhibitor of pathogen microbes and it may convert into probiotic bacteria (Kerry et al., 2018).
Probiotics are beneficial bacteria; they alter the intestinal microflora balance and inhibit the growth of pathogen microbes. To perform beneficial effects, probiotics must survive in the gastrointestinal tract, persist in the host and prove safety for consumers. To survive in the gut, the organism must be tolerant of low pH and bile toxicity prevalent in the upper digestive tract (Shokryazdan et al., 2016). A study had successfully isolated LAB from 144 kinds of plara (a fermented fish made in Thailand), namely A. viridans, E. avium, E. faecalis, E. faecium, E. hirae, E. thailandicus, L. plantarum, L. lactis, L. paracasei, P. pentosaceus, P. acidilactici, T. halophilus, W. cibaria, W. confusa, W. paramesenteroides, and W. viridescens (Miyashita et al., 2012).
Studies on rabbit meat bekasam are currently non-existent in Indonesia. Therefore, this study investigated the chemical and microbiological characteristics of rabbit meat bekasam (during the fermentation process) in order to isolate, characterize (in vitro and in vivo), and identify LAB as the candidate probiotic bacteria. The isolated bacteria can be used as the starter for rabbit meat bekasam.
Materials and Methods
Nine healthy New Zealand White crossbreed aged 3 months old weighing 1,800±53 g were purchased from Rajawali Farm, Sumedang, Indonesia. The rabbits were slaughtered used halal method according to Fuseini et al. (2017). Carcas was obtained by removing the blood, skin, distal portions of legs, distal part of the tail, organs located in the thorax and neck (lungs, oesophagus, trachea, thymus, and heart), genital organs, urinary bladder, gastrointestinal tract, liver, and kidneys. After deboning, the whole meat was chopped and rationed to seven batches for seven-day observation. Rabbit meat bekasam was prepared with a method by Sari et al. (2018) with a slight modification. Rabbit meat was marinated with 10% salt for 6 h and stored in a sterile container along with Setra Ramos® local rice (1:1 ratio). The container was tightly closed and incubated at room temperature (27.0±2.0°C) for 7 d.
The proximate composition of rabbit meat bekasam was measured according to the methods of AOAC International (2012). The moisture content of weight loss was calculated after 12 h oven-drying at 105°C (Digital drying oven DOD-150, Raypa, Barcelona, Spain). The protein content was measured using an automatic Kjeldahl nitrogen analyzer (AutoKjeldahl Unit K-370, Büchi Labortechnik, Flawil, Switzerland). Fat content was determined with the Soxhlet method using a solvent extraction system (SoxtecTM 2050 automated analyzer, FOSS Analytical, Hillerød, Denmark). A dry ashing method to determine ash content was conducted by incinerating the meat samples in a furnace (Thermolyne FD1410M, ThermoFisher Scientific, Waltham, MA, USA) at 550°C. Total carbohydrate was calculated (by difference) using the formula: Total CHO = 100 – (moisture% + fat% + protein% + ash%).
The pH value of rabbit meat bekasam was measured using a pH meter (3510 Advanced Bench pH Meters, Jenway, Staffordshire, UK). Five gram rabbit meat bekasam was blended with 20 mL distilled water in a homogenizer for 60 s (Ultra-Turrax T25, IKA, Darmstadt, Germany). Lactic acid content was determined using the standard titration procedure for total titratable acidity.
Twenty five grams bekasam sample from each treatment was transferred to 50 mL of sterile saline solution and homogenized for 90 s, and serial dilutions were prepared by mixing 1 mL of the homogenized sample with 9 mL of sterile saline solution. Total bacteria and total LAB were enumerated by plating samples on Nutrient Agar (NA, M001, HiMedia) and Lactobacillus MRS Agar (MRSA, M6411, HiMedia), respectively after aerobic incubation at 37°C for 24 h. Total yeast and moulds were count by plating serial dilution on Malt Extract Agar (MEA, M137, HiMedia) and Potato Dextrose Agar (PDA, M137, HiMedia), respectively, after aerobic incubation at 27°C for 48 h. Total Coliform was count on MacConkey Agar (MCA, MH081, HiMedia) media after incubation at 37°C for 48 h. The formed colonies were count and expressed as colony forming units of the suspension (CFU/g).
LAB was isolated by suspending 25 g sample in 225 mL of Lactobacillus MRS Broth (MRSB, M369, HiMedia) followed by anaerobic incubation at 37°C for 24 h for enrichment, and the MRSB was serially diluted with sterile saline solution. Appropriate dilutions were spread on MRSA plates containing 0.3% (w/v) CaCO3 (Calcium Carbonate, GRM1044, HiMedia) for 48-hour incubation at 37°C. LAB produces lactic acid and reacts with CaCO3 to produce soluble lactate calcium, characterized by a clear zone around the growing bacterial colonies. The possible identification of LAB colonies was tested for catalase activity, motility, and Gram staining. All catalase-negative, non-motil and positive Gram staining colonies were streaked on the MRSA and incubated at 37°C for 48 h to obtain pure colonies. They were stored in slanted semi-solid MRSA at 4°C.
The viability in low pH (pH 2.0 and pH 3.0) were examined using 0.1 N HCl to manipulate the MRSB medium. One mL of 24-h-old LAB culture was pipetted into 9 mL of MRSB pH 2.0 and pH 3.0, incubated at 37°C for 3 h. Before and after a 3-hour incubation, total cells were enumerated in MRSA pouring method, followed by a 48-hour incubation at 37°C (Ngom, 2000). To measure bacteria survival against bile salt, 1 mL of 24-h-old LAB culture was pipetted into 9 mL of MRSB that contained 0.5% and 1% bile salt (Bile salt, RM008, HiMedia), incubated at 37°C for 48 h, and count the cell using MRSA pouring method.
Antimicrobial activity was determined by agar well diffusion assay according to Tagg et al. (1976). The pathogens in this study were Gram-positive bacteria (Staphylococcus aureus ATCC 6538; Listeria monocytogenes ATCC 7644) and Gram-negative bacteria (Escherichia coli ATCC 11229; Salmonella thypimurium ATCC 14088) obtained from the Central Laboratory of Padjadjaran University, Jatinangor, Indonesia. Standard chloramphenicol 30 mg/mL (Chloramphenicol, TC204, HiMedia) were used as a positive control. The pathogens cultured in Nutrient Broth (NB, M002, HiMedia) were spread on the NA plate 5 mm diameter wells were made in the plate, poured with 40 μL of LAB isolate, and incubated at 37°C for 24 h. The diameter of the clear zone formed around the well after incubation was measured to confirm the antimicrobial activity.
LAB isolates were cultured in MRSB (pH 7.0) for 1d and bacterial cells were collected by centrifugation at 3,185×g for 10 min (Sigma 1-16K, Sigma-Aldrich, Osterode am Harz, Germany). The genomic DNA was extracted in a method by Zhu et al. (1993) with modification by Mustopa and Fatimah (2014). The pellet was resuspended with TE buffer (10 mM Tris-HCl pH 8.0, 1 mM EDTA), 40 μL of lysozyme (60 mg/mL; Lysozyme, PC0710, Vivantis), incubated at 37 °C for 60 min and added with 200 μL 10% sodium dodecyl sulfate, 100 μL 5 M NaCl, and 80 μL 10% centrimide. Furthermore, the solution was warmed at 68°C for 30 min, added with equal amount of chloroform, and proceed with centrifugation at 14,953×g speed for 10 min. The supernatant was harvested and 1 mL of ethanol was added. The mixture was shaken again and then centrifuged at 14,953×g for 10 min. after being air-dried, the DNA was dissolved in TE buffer and the concentration was adjusted to 10 μg/mL DNA and stored at −20°C for further analysis.
For 16S rRNA sequencing, primers 8F (5'-AGA GTT TGA TCA TGG CTC AG-3'; positions 8 to 27 bp) and 15R (5'-AAGGAG GTG ATC CAA CCG CA-3'; positions 1,541 to 1,522 bp) (Invitrogen, ThermoFisher Scienctific) were used to amplify the full length of bacterial 16S rRNA fragment (Chao et al., 2008). Each 25 μL polymerase chain reaction (PCR) mixture (Invitrogen, 10572014, ThermoFisher Scienctific) contained 10 mM Tris L-HCl (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, 200 μM of each dNTP, 400 nM of each primer, 1 U of Taq polymerase, and 10 ng of the DNA template. The PCR was operated at 96°C for 5 min, and performed 35 cycles consisting of 96°C for 1 min, 58°C for 3 min, 72°C for 1 min; and 72°C for 7 min. The PCR products were subjected to gel electrophoresis in 1% agarose gel, followed by ethidium bromide staining.
The DNA sequencing was performed in Macrogen, Korea. An online BLAST analysis performed similarity searches with sequences were performed by an online BLAST analysis in the National Centre for Biotechnology Information (NCBI). For the phylogenetic analysis, sequences were aligned by using the CLUSTAL X software (Thompson et al., 1997), while the phylogenetic tree was constructed by the neighbor-joining method (Saitou and Nei, 1987).
Lactobacillus buchneri was cultured in MRSB, incubated at 37°C for 24 h, and centrifuged 8,848×g for 10 min at 4°C. Supernatants were discarded, and cell pellets were washed three times with deionized water. For cell suspension preparation, sterile saline was used as a diluent.
The study was performed on 20 male BALB/c mice aged 6–8 weeks old, weighing about 22–32 g, purchased from PT Biofarma (Bandung, Indonesia). Experimental animal handling was following the Polish law on the protection of animals. The Ethics Committee of Universitas Padjadjaran approved all experiments with Ethical Committee No. 690/UN6.KEP/ EC/2019. The mice were housed (three per cage) and maintained under conventional conditions (room temperature 22±2°C, 12 h day/night cycle). The standard diet contained 13% maximum water content, 10%–12% protein, 5% fat, 8% fibre, 14% ash, 3% calcium, and 0.7% phosphor.
The mice were distributed into five dietary groups (four replicates each). After seven days of acclimatization and fasting for 16 h, each group was administered (by oral gavage) a single dose of one of five treatments. The treatments included P1: 0.2 mL sterile saline (negative control); P2: 0.2 mL 1×108 CFU of Bifidobacterium bifidum in 1 mL sterile saline (positive control); P3: 0.2 mL 1×107 CFU/mL of L. buchneri in 1 mL sterile saline; P4: 0.2 mL 1×108 CFU/mL of L. buchneri in 1 mL sterile saline; and P5: 0.2 mL 1×109 CFU/mL of L. buchneri in 1 mL sterile saline. The mice were fed for seven consecutive days. On the eighth day, each animal was inoculated with a single 0.2 mL dose of 1×107 CFU/mL of Salmonella typhimurium in sterile saline. After three days, the mice were sacrificed by cervical dislocation and were anesthetized by 100 mg/kg (w/w) intraperitoneal injection of ketamine (Ketamil, Iliuum, New South Wales, Australia) and 20 mg/kg (w/w) xylazine (Xyla, Interchemie, AC Castenray, Netherlands).
Fresh mice feces were collected and pooled daily from each group from day 0 to day 4 to enumerate the LAB, Coliform, and Salmonella using the spread plate method. 1 g fecal was suspended briefly in 9 mL phosphate-buffered saline (PBS) and vortexed for 1 min. The samples were serially diluted in sterile diluents, and 100 μL 10−4 to 10−6 dilutions were streaked on different media for bacterial count, MRSA for total LAB, MacConkey agar for coliform, and Xylose Lysine Deoxycholate agar (XLDA, M031, HiMedia) for Salmonella. After 24 h incubation at 37°C, the colonies on the plates were counted and the microbial population was expressed as CFU/g.
Results and Discussion
The proximate composition of rabbit meat bekasam during fermentation is presented in Table 1. The moisture content of bekasam was significantly decreased (p<0.05) from 66.50% to 55.05%. Salt which was added on bekasam have the ability to decrease the moisture content. According to Fennema (1996), salt can decrease moisture content is due to the ability of sodium and chloride ions to associate with water molecules. Similarly, Majumdar and Basu (2010) reported that the water content of fermented fish product (Lona ilish) from India was decreased during fermentation.
The fat content of bekasam was significantly decreased (p<0.05) during fermentation procces. The bacteria and yeast performed lipolytic activity which hydrolyzed fat molecules into free fatty acids, hence, reducing the fat content. It was in line with Desniar et al. (2009) on the fermentation process of chub mackerel (Rastrelliger sp.).
The carbohydrate content of bekasam was reduced significantly (p<0.05). bekasam as a rice-based fermentation product was degraded by yeast and mould that produced an extracellular amylolytic enzymes (α-amylase and glucoamylase). Kumar et al. (2013) stated that starch was hydrolyzed into maltose and glucose by amylolytic enzyme which then converted into lactid acid and other organic acid, decreasing the carbohydrate content during fermentation. It was confirmed by the decreasing pH and the increasing lactid acid content in bekasam during fermentation in this research. On the other hand, protein content was increased significantly (p<0.05). Protease was hydrolyzing protein into polypeptide and amino acid which was used by microbes to increase microbial cell. Microbial cells are mostly build on protein; hence, microbes are known as Single Cell Protein (SCP) or natural protein concentrates (Kurbanoğlu, 2001). It was reflected from the increasing LAB and yeast in this study (Table 2).
There was no significant difference in the ash content (p>0.05). It indicated that during the fermentation process, the microbe used and produced mineral in a small amount. The minerals of the rabbit meat included potassium, phosphor, sodium, magnesium, calcium, zinc, iron, copper, and manganese (Hermida et al., 2006).
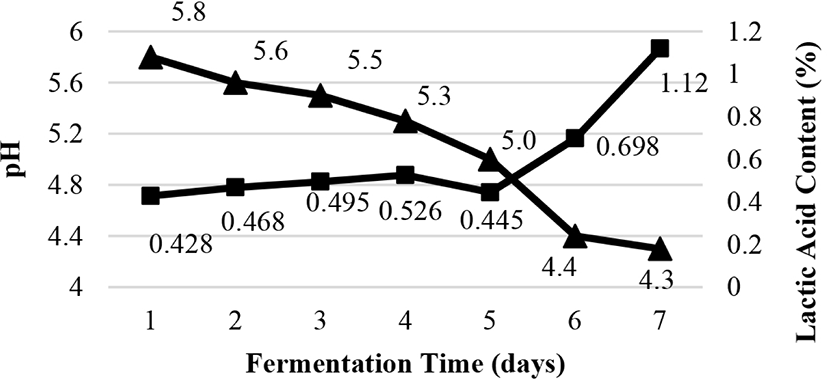
Titratable acidity and pH are illustrated in Fig. 1. The pH was reduced from 5.8 to 4.3, whereas the titratable acidity was increased from 0.428% to 1.12%. It was due to the accumulation of lactic acid and organic acid generated during the fermentation process which promoted the decrease in pH value. The growth of LAB converted carbohydrates to lactic, acetic, formic, caproic, propionic, butyric, and valeric acids (Zalán et al., 2010). Our findings are in agreement to previous studies on lower pH values of fermented fish during fermentation (Desniar et al., 2012; Paludan-Muller et al., 2002).
Microbiological characteristics of bekasam during fermentation are shown in Table 2. The total bacteria was improved across 6 days of fermentation and decreased on the 7th day because low pH (4.4) was not suitable for the growth of several bacteria. Total LAB and yeast was increased from 3.82×106 to 4.67×108 CFU/g, and 9.89×106 to 3.42×108 CFU/g, respectively. In this study, total yeast was linear to total LAB during fermentation. LAB generates organic acids that may reduce pH and potentially stimulates the growth of yeast. Meanwhile, yeast produces vitamin and amino acid to support LAB growth (Fleet, 1990). Paludan-Muller et al. (2002) reported that isolated LAB and yeast were the dominant fermenting microorganisms for plaa-som, the fermented fish produced in Thailand.
The total mould declined from 4.86×103 to 4.34×101 CFU/g which may due to high salinity in fermentation. Coliform was present until the second day of fermentation because it could not survive in high salinity. Salt render microbial cells to undergo osmotic shock, resulting in the loss of water from the cell, and subsequently, cell death or retarded growth (Anihouvi et al., 2012). Total Coliform in this study was similar to <10 CFU/g in fermented fish (Hout-Kasef) produced in Saudi Arabia (Gassem, 2019).
LAB isolation using MRSA with calcium carbonate produced 9 isolates. The colony morfology of isolated bacteria illustrated in Table 3 includes Gram-positive, catalase-negative, and non-motile. It indicated that indicating all bekasam isolates were corresponded to the LAB criteria proposed by Kerry et al. (2018).
Resistance to low pH and bile salts are the key factors to predict the survival and growth of potential probiotic strains in gastrointestinal conditions. According to Sahadeva et al. (2011), low pH incubation was viable at pH 2.0 and pH 3.0 for 3 h as it stimulates bacterial residency. The viability of LAB in low pH and bile salt are as presented in Table 4 and Table 5 respectively. In general, total LAB counts of the nine isolates declined after exposure to low pH, and they were tolerant in pH 3.0 compared to pH 2.0. The acidity tolerance of LAB is attributed to a constant gradient between extracellular and cytoplasmic pH. When internal pH reached the threshold, cellular functions were inhibited and the cells died (Kashket, 1987).
Total LAB declined after exposure to 0.5% bile salts, and four LAB isolates (3.1, 5.1, 5.2, and 6.3) could not grow after exposure to 1.5% bile salts. Exposure to bile salts triggered the disruptions of cellular homeostasis which dissociated lipid bilayer and integral protein from their cell membranes, causing leakage in bacterial content and finally, cell death. (Tokatli et al., 2015). Isolate 6.1 and 7.1 in this study were highly tolerant to low pH and bile salt, producing 106 CFU/mL total LAB which was the minimum total probiotic microbes (Sahadeva et al., 2011).
The antimicrobial activity of all LAB isolates is presented in Table 6. All isolates exhibited a broad spectrum antimicrobial active against Gram-positive and Gram-negative pathogenic bacteria. It was in line with Sari et al. (2018) who examined the antimicrobial activities of fermented fish and discovered that LAB isolates might inhibit Gram-positive (S. aureus ATCC 25923 with 12.7 mm inhibition zone) and Gram-negative (Salmonella sp. with 7.3 mm) pathogenic bacteria. The activity of isolate 6.1 was highest against S. aureus with a 16.4 mm inhibition zone, and isolate 3.1 was the least for E. coli (6.2 mm).
LAB isolates can inhibit the growth of pathogenic microbes because LAB produces antimicrobial compounds during the fermentation process, such as organic acid (lactic acid and acetic acid), diacetyl, ethanol, hydrogen peroxide, reuterin, acetaldehyde, acetoin, carbon dioxide, and bacteriocins (García-Cano et al., 2014). This study indicated that isolate 6.1 exhibited the highest antibacterial activity against all pathogens tested except L.monocytogenes.
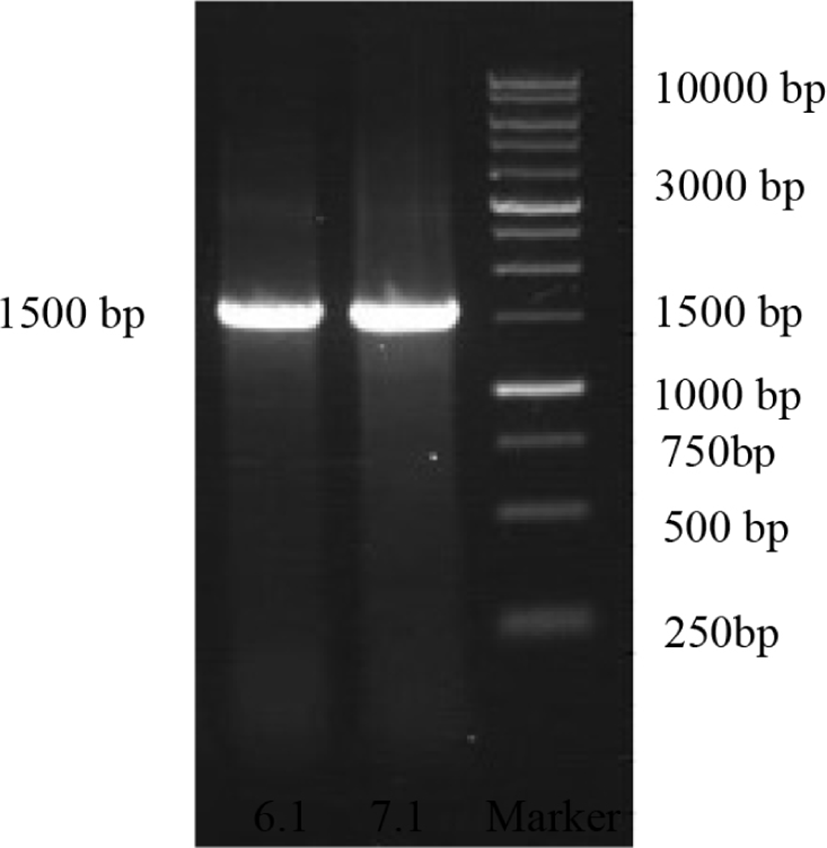
Probiotic candidate test on the viability of pH, bile salt, and antimicrobial activity concluded that isolates 6.1 and 7.1 were the best probiotic candidate isolates. Both isolates were subject to molecular identification using 16S rRNA gene analysis. The result of the 16S rRNA gene amplification could be identified from any fragment of PCR product with the size of 1,500 base pairs (bp) as the desired measurement (Fig. 2).
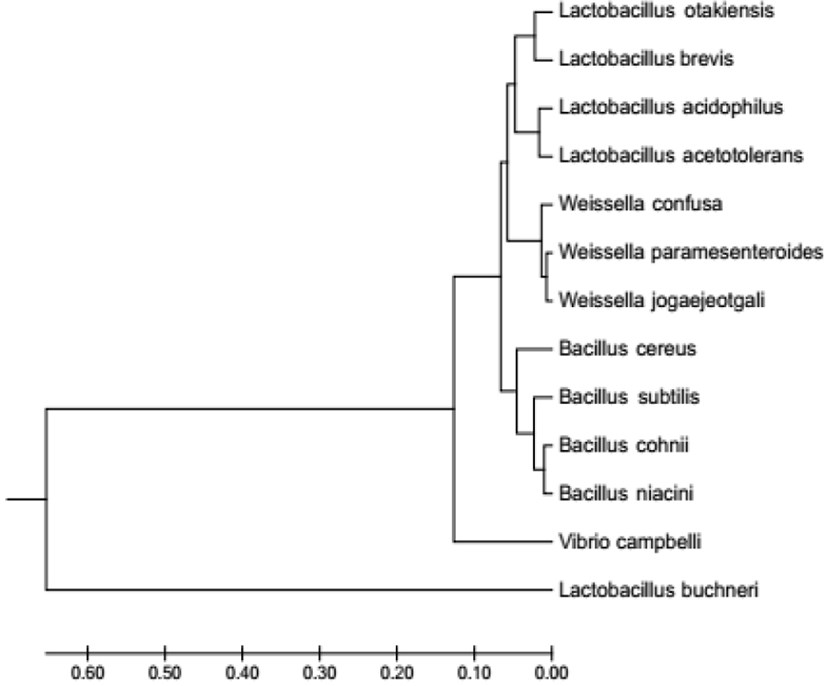
The homology analysis based on BLAST and SIM revealed that isolate 6.1 had a genetic relationship with Lactobacillus buchneri, and isolate 7.1 had the closest genetic relationship with Weisella paramesentoroides with 99% homology level. According to Clarridge (2004), the similarity level of a species is 94%. Phylogenetic tree of isolates 6.1 and 7.1 based on gene sequence of 16S rRNA is shown in Fig. 3.
One of the identified isolates (L. buchneri) was selected as a single starter for bekasam future production because it acquired the Generally Recognized As Safe (GRAS) status (FAO/WHO, 2002). According to Fessard and Remize (2017), Weisella spp. was not a GRAS starter.
L. buchneri probiotic isolates were studied in vivo using BALB/c strain mice to identify the safety use in fermented food. Mice body weight was recorded (Table 7) to indicate the adverse effect of substrate in animal study. Table 7 showed that mice body weight increased during L. buchneri treatment, and maintained after infection with S. typhimurium. It indicated that L. buchneri positifely affected mice’s immune system. It was in line with El-Jakee et al. (2010) and Shokryazdan et al. (2016) that mice receiving probiotics did not undergo body weight loss.
| Treatment | Body weight (g) | ||
|---|---|---|---|
| Day 1 | Day 7 | Day 12 | |
| P1 | 22.6±0.9 | 24.5±0.5 | 25.1±1.3 |
| P2 | 22.5±0.7 | 24.8±0.8 | 26.2±1.0 |
| P3 | 20.5±1.4 | 22.7±1.2 | 24.5±0.7 |
| P4 | 21.5±1.0 | 23.1±1.3 | 24.6±0.9 |
| P5 | 22.8±1.2 | 24.5±1.2 | 25.6±1.2 |
P1, 0.2 mL of sterile saline (negative control); P2, 0.2 mL dose of 1×108 CFU of Bifidobacterium bifidum in 1 mL sterile saline (positive control); P3, 0.2 mL dose of 1×107 CFU/mL of Lactobacillus buchneri in 1 mL sterile saline; P4, 0.2 mL dose of 1×108 CFU/mL of L. buchneri in 1 mL sterile saline; P5, 0.2 mL dose of 1×109 CFU/mL of L. buchneri in 1 mL sterile saline.
The results of microbiological test in mice feces are shown in Fig. 4. Results revealed that total fecal LAB population in the negative control (P1) was lower throughout the observation days. Total LAB declined after the third day of S. typhimurium infection in mice which was administered with probiotic treatment (P2, P3, and P4). L. buchneri cells treatment increased total LAB in mice feces on the seventh day of L. buchneri administration, and decreased after the L. buchneri administration stopped and S. typhimurium infection started. Accordingly, L. buchneri could inhibit S. typhimurium infection by competing for essential nutrition; therefore, total LAB decreased after the L. buchneri administration stopped.
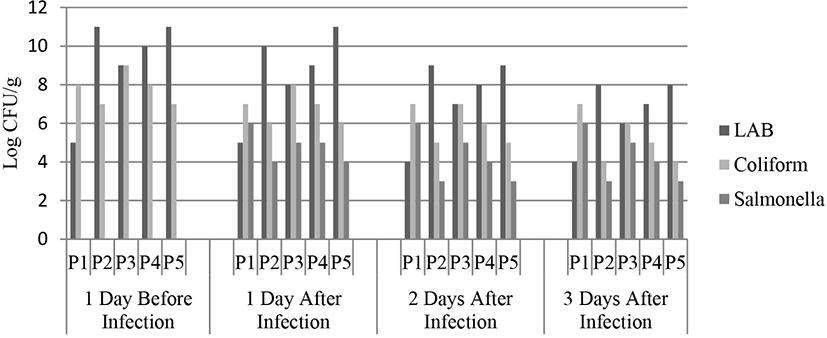
Total Coliform and Salmonella of mice feces in probiotic treatment were decreased. Salmonella was non-existent in mice feces on the day before infection. It indicated that the mice were healthy and not infected by Salmonella. The effect of probiotic administration on total coliform may increase due to competition between bacteria to adhere to the epithelial cells of the intestine Sherman et al. (2005). Previous studies reported the declining total intestinal Salmonella due to probiotics effect (El-Jakee et al., 2010; Vasilica and Balotescu, 2006).
Conclusion
Rabbit meat bekasam is considered a nutritional and health-promoting food. This study isolated Lactobacillus buchneri from rabbit meat bekasam and further analysis showed that it was resistant to acid pH and bile salts in vitro, and exhibited in vitro antimicrobial activity against Gram-positive and Gram-negative pathogen bacteria. The in vivo analysis demonstrated that L. buchneri was safe up to 109 CFU/g in a 7-day treatment period. L. buchneri could reduce the populations of harmful intestinal bacteria and pathogenic bacteria while increasing beneficial bacterial populations. No adverse effects were observed on the growth of the experimental animals.













